Integrate single-cell RNA-seq data using various integration methods.
For ChromatinAssay, the current workflow uses TFIDF + SVD/LSI preprocessing.
In this setting, Uncorrected is supported directly and Harmony5 will be
automatically redirected to the legacy Harmony workflow. Seurat and RPCA
are currently not supported for ChromatinAssay.
Usage
integration_scop(
srt_merge = NULL,
batch,
append = TRUE,
srt_list = NULL,
assay = NULL,
integration_method = c("Uncorrected", "Seurat", "CCA", "RPCA", "scVI", "PeakVI",
"PoissonVI", "WNN", "MultiMAP", "GLUE", "scVI5", "MNN", "fastMNN", "fastMNN5",
"Harmony", "Harmony5", "Scanorama", "BBKNN", "CSS", "Coralysis", "LIGER", "Conos",
"ComBat"),
compute_lisi = FALSE,
lisi_label_colnames = NULL,
lisi_reduction = NULL,
lisi_dims = NULL,
lisi_prefix = NULL,
lisi_tool_name = NULL,
lisi_perplexity = 30,
lisi_nn_method = c("auto", "exact"),
lisi_tol = 1e-05,
lisi_max_iter = 50,
compute_metrics = FALSE,
metrics_batch_col = NULL,
metrics_celltype_col = NULL,
metrics_reduction = NULL,
metrics_cluster_col = NULL,
metrics_tool_name = NULL,
metrics_k_graph = 15,
do_normalization = NULL,
normalization_method = "LogNormalize",
do_HVF_finding = TRUE,
HVF_source = "separate",
HVF_method = "vst",
nHVF = 2000,
HVF_min_intersection = 1,
HVF = NULL,
do_scaling = TRUE,
vars_to_regress = NULL,
regression_model = "linear",
scale_within_batch = FALSE,
linear_reduction = "pca",
linear_reduction_dims = 50,
linear_reduction_dims_use = NULL,
linear_reduction_params = list(),
force_linear_reduction = FALSE,
nonlinear_reduction = "umap",
nonlinear_reduction_dims = c(2, 3),
nonlinear_reduction_params = list(),
force_nonlinear_reduction = TRUE,
neighbor_metric = "euclidean",
neighbor_k = 20L,
cluster_algorithm = "louvain",
cluster_resolution = 0.6,
seed = 11,
verbose = TRUE,
...
)Arguments
- srt_merge
A merged `Seurat` object that includes the batch information.
- batch
A character string specifying the batch variable name.
- append
Whether the integrated data will be appended to the original Seurat object (
srt_merge). Default isTRUE.- srt_list
A list of
Seuratobjects to be checked and preprocessed.- assay
Which assay to use. If
NULL, the default assay of the Seurat object will be used. When the object also containsChromatinAssay, the default assay and additionalChromatinAssaywill be preprocessed sequentially.- integration_method
A character vector specifying the integration method to use. Supported methods are:
"Uncorrected","Seurat","CCA","RPCA","scVI","PeakVI","PoissonVI","WNN","MultiMAP","GLUE","scVI5","MNN","fastMNN","fastMNN5","Harmony","Harmony5","Scanorama","BBKNN","CSS","Coralysis","LIGER","Conos","ComBat". Default is"Uncorrected". ForChromatinAssay, prefer"Uncorrected"or"Harmony5"; the latter is automatically switched to"Harmony".- compute_lisi
Whether to compute LISI scores on the integrated result. Default is
FALSE.- lisi_label_colnames
Character vector of metadata columns used to compute LISI. If
NULLandcompute_lisi = TRUE,batchwill be used when it is a single metadata column name.- lisi_reduction
Dimensional reduction used for LISI computation. Default is
NULL, which usesDefaultReduction()from the integrated object.- lisi_dims
Dimensions used from
lisi_reduction. Default isNULL, which uses all available dimensions.- lisi_prefix
Prefix used when storing LISI metadata columns. Default is
NULL, which useslisi_reduction.- lisi_tool_name
Name of the tool entry used to store LISI results. Default is
NULL, which usespaste0(lisi_prefix, "_LISI").- lisi_perplexity
Effective neighborhood size used by LISI. Default is
30.- lisi_nn_method
Nearest-neighbor backend passed to
RunLISI(). One of"auto"or"exact". Default is"auto".- lisi_tol
Tolerance used in the LISI binary search. Default is
1e-5.- lisi_max_iter
Maximum iterations used in the LISI binary search. Default is
50.- compute_metrics
Whether to compute integration summary metrics on the selected reduction. Default is
FALSE.- metrics_batch_col
Metadata column used for batch-mixing metrics. Default is
NULL, which usesbatchwhen it is a single metadata column name.- metrics_celltype_col
Metadata column used for biological conservation metrics. Default is
NULL.- metrics_reduction
Reduction used for integration metric computation. Default is
NULL, which usesDefaultReduction()from the integrated object.- metrics_cluster_col
Metadata column used as cluster labels for
celltype_NMI,celltype_ARI, andcelltype_purity. Default isNULL, which resolves the integrated cluster column automatically when possible.- metrics_tool_name
Name of the tool entry used to store integration metrics. Default is
NULL, which usespaste0(integration_method, "_metrics").- metrics_k_graph
Number of neighbors used for graph-connectivity computation. Default is
15.- do_normalization
Whether data normalization should be performed. Default is
TRUE.- normalization_method
The normalization method to be used. Possible values are
"LogNormalize","SCT","TFIDF", and"scran". Default is"LogNormalize".- do_HVF_finding
Whether to perform high variable feature finding. If
TRUE, the function will force to find the highly variable features (HVF) using the specified HVF method.- HVF_source
The source of highly variable features. Possible values are
"global"and"separate". Default is"separate".- HVF_method
The method to use for finding highly variable features. Options are
"vst","mvp","disp", or"scran". Default is"vst".- nHVF
The number of highly variable features to select. If NULL, all highly variable features will be used. Default is
2000.- HVF_min_intersection
The feature needs to be present in batches for a minimum number of times in order to be considered as highly variable. Default is
1.- HVF
A vector of feature names to use as highly variable features. If NULL, the function will use the highly variable features identified by the HVF method.
- do_scaling
Whether to perform scaling. If
TRUE, the function will force to scale the data using the Seurat::ScaleData function.- vars_to_regress
A vector of variable names to include as additional regression variables. Default is
NULL.- regression_model
The regression model to use for scaling. Options are
"linear","poisson", or"negativebinomial". Default is"linear".- scale_within_batch
Whether to scale data within each batch. Only valid when the
integration_methodis one of"Uncorrected","Seurat","MNN","Harmony","BBKNN","CSS","ComBat".- linear_reduction
The linear dimensionality reduction method to use. Options are
"pca","svd","ica","nmf","mds", or"glmpca". Default is"pca".- linear_reduction_dims
The number of dimensions to keep after linear dimensionality reduction. Default is
50.- linear_reduction_dims_use
The dimensions to use for downstream analysis. If
NULL, estimated dimensions stored in the linear reduction will be used when available; otherwise, the first up to50dimensions will be used as a fallback.- linear_reduction_params
A list of parameters to pass to the linear dimensionality reduction method.
- force_linear_reduction
Whether to force linear dimensionality reduction even if the specified reduction is already present in the Seurat object.
- nonlinear_reduction
The nonlinear dimensionality reduction method to use. Options are
"umap","umap-naive","tsne","dm","phate","pacmap","trimap","largevis", or"fr". Default is"umap".- nonlinear_reduction_dims
The number of dimensions to keep after nonlinear dimensionality reduction. If a vector is provided, different numbers of dimensions can be specified for each method. Default is
c(2, 3).- nonlinear_reduction_params
A list of parameters to pass to the nonlinear dimensionality reduction method.
- force_nonlinear_reduction
Whether to force nonlinear dimensionality reduction even if the specified reduction is already present in the Seurat object. Default is
TRUE.- neighbor_metric
The distance metric to use for finding neighbors. Options are
"euclidean","cosine","manhattan", or"hamming". Default is"euclidean".- neighbor_k
The number of nearest neighbors to use for finding neighbors. Default is
20.- cluster_algorithm
The clustering algorithm to use. Options are
"louvain","slm", or"leiden". Default is"louvain".- cluster_resolution
The resolution parameter to use for clustering. Larger values result in fewer clusters. Default is
0.6.- seed
Random seed for reproducibility. Default is
11.- verbose
Whether to print the message. Default is
TRUE.- ...
Additional arguments to be passed to the integration method functions.
Value
A Seurat object.
For ChromatinAssay, integrated outputs are additionally normalized to the
ATAC naming convention used in scop, including *lsi, *UMAP2D, cluster
aliases, and ATAC_default_* metadata when available.
Examples
data(panc8_sub)
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "Uncorrected",
nHVF = 500,
linear_reduction_dims = 20,
linear_reduction_dims_use = 1:10,
nonlinear_reduction_dims = 2,
compute_lisi = TRUE,
lisi_label_colnames = "tech",
lisi_perplexity = 10
)
#> ◌ [2026-05-14 07:53:32] Run integration workflow...
#> ℹ [2026-05-14 07:53:33] Split `srt_merge` into `srt_list` by "tech"
#> ℹ [2026-05-14 07:53:34] Checking a list of <Seurat>...
#> ! [2026-05-14 07:53:34] Data 1/5 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:53:34] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:36] Perform `Seurat::FindVariableFeatures()` on 1/5 of `srt_list`...
#> ! [2026-05-14 07:53:36] Data 2/5 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:53:36] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 2/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:38] Perform `Seurat::FindVariableFeatures()` on 2/5 of `srt_list`...
#> ! [2026-05-14 07:53:38] Data 3/5 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:53:38] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 3/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:40] Perform `Seurat::FindVariableFeatures()` on 3/5 of `srt_list`...
#> ! [2026-05-14 07:53:40] Data 4/5 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:53:40] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 4/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:42] Perform `Seurat::FindVariableFeatures()` on 4/5 of `srt_list`...
#> ! [2026-05-14 07:53:42] Data 5/5 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:53:42] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:44] Perform `Seurat::FindVariableFeatures()` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:53:44] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:53:45] Number of available HVF: 500
#> ℹ [2026-05-14 07:53:45] Finished check
#> ℹ [2026-05-14 07:53:47] Perform Uncorrected integration
#> Warning: Layer ‘scale.data’ is empty
#> ℹ [2026-05-14 07:53:47] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 07:53:47] Perform "pca" linear dimension reduction
#> ℹ [2026-05-14 07:53:48] Adjust neighbor k from 20 to 20 for small-sample clustering
#> ℹ [2026-05-14 07:53:48] Perform `Seurat::FindClusters()` with "louvain"
#> ℹ [2026-05-14 07:53:48] Reorder clusters...
#> ℹ [2026-05-14 07:53:49] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 07:53:49] Perform umap nonlinear dimension reduction using Uncorrectedpca (1:10)
#> ℹ [2026-05-14 07:53:55] Perform umap nonlinear dimension reduction using Uncorrectedpca (1:10)
#> ℹ [2026-05-14 07:54:02] Compute LISI scores from reduction "UncorrectedpcaUMAP2D"
#> ℹ [2026-05-14 07:54:02] Compute LISI scores from reduction "UncorrectedUMAP2D"
#> ✔ [2026-05-14 07:54:02] Stored LISI scores in metadata: "UncorrectedpcaUMAP2D_tech_LISI" and "UncorrectedUMAP2D_tech_LISI"
#> ✔ [2026-05-14 07:54:05] Uncorrected integration completed
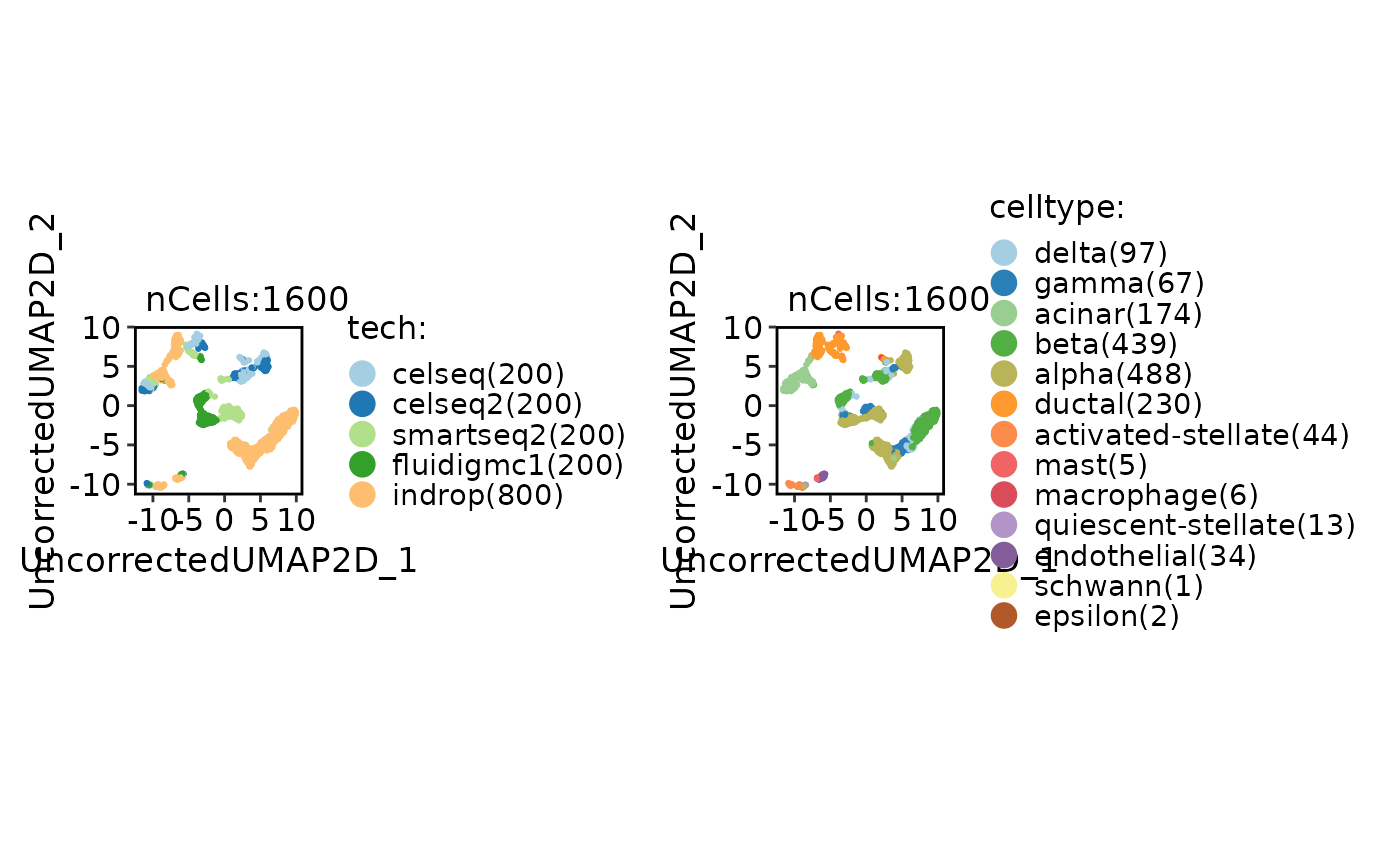
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "UncorrectedUMAP2D"
)
 LISIPlot(
panc8_sub,
features = c("UncorrectedpcaUMAP2D_tech_LISI", "UncorrectedUMAP2D_tech_LISI")
)
LISIPlot(
panc8_sub,
features = c("UncorrectedpcaUMAP2D_tech_LISI", "UncorrectedUMAP2D_tech_LISI")
)
 panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "LIGER"
)
#> ◌ [2026-05-14 07:54:06] Run integration workflow...
#> ℹ [2026-05-14 07:54:06] Split `srt_merge` into `srt_list` by "tech"
#> ℹ [2026-05-14 07:54:07] Checking a list of <Seurat>...
#> ℹ [2026-05-14 07:54:07] Data 1/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:07] Perform `Seurat::FindVariableFeatures()` on 1/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:08] Data 2/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:08] Perform `Seurat::FindVariableFeatures()` on 2/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:08] Data 3/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:08] Perform `Seurat::FindVariableFeatures()` on 3/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:09] Data 4/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:09] Perform `Seurat::FindVariableFeatures()` on 4/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:09] Data 5/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:09] Perform `Seurat::FindVariableFeatures()` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:10] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:54:10] Number of available HVF: 2000
#> ℹ [2026-05-14 07:54:10] Finished check
#> Warning: Layer ‘ligerScaleData’ is empty
#> ℹ [2026-05-14 07:54:16] Prepare rliger layer "ligerScaleData" ...
#> ℹ [2026-05-14 07:54:17] Perform LIGER integration
#> ℹ [2026-05-14 07:54:23] Adjust neighbor k from 20 to 20 for small-sample clustering
#> ℹ [2026-05-14 07:54:23] Perform `Seurat::FindClusters()` with "louvain"
#> ℹ [2026-05-14 07:54:23] Reorder clusters...
#> ℹ [2026-05-14 07:54:24] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 07:54:24] Perform umap nonlinear dimension reduction using LIGER (1:20)
#> ℹ [2026-05-14 07:54:31] Perform umap nonlinear dimension reduction using LIGER (1:20)
#> ℹ [2026-05-14 07:54:37] Perform umap nonlinear dimension reduction using Uncorrectedpca (1:20)
#> ✔ [2026-05-14 07:54:46] LIGER integration completed
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "Harmony",
compute_lisi = TRUE,
lisi_label_colnames = "tech"
)
#> ◌ [2026-05-14 07:54:46] Run integration workflow...
#> ℹ [2026-05-14 07:54:46] Split `srt_merge` into `srt_list` by "tech"
#> ℹ [2026-05-14 07:54:47] Checking a list of <Seurat>...
#> ℹ [2026-05-14 07:54:48] Data 1/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:48] Perform `Seurat::FindVariableFeatures()` on 1/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:48] Data 2/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:48] Perform `Seurat::FindVariableFeatures()` on 2/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:49] Data 3/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:49] Perform `Seurat::FindVariableFeatures()` on 3/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:49] Data 4/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:49] Perform `Seurat::FindVariableFeatures()` on 4/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:50] Data 5/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:50] Perform `Seurat::FindVariableFeatures()` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:50] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:54:51] Number of available HVF: 2000
#> ℹ [2026-05-14 07:54:51] Finished check
#> Warning: Layer ‘scale.data’ is empty
#> ℹ [2026-05-14 07:55:16] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 07:55:16] Perform linear dimension reduction("pca")
#> ℹ [2026-05-14 07:55:17] Perform Harmony integration
#> ℹ [2026-05-14 07:55:17] Using "Harmonypca" (1:20) as input
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 't': ‘Z_corr’ is not a valid field or method name for reference class “Rcpp_harmony”
LISIPlot(
panc8_sub,
features = c("HarmonypcaUMAP2D_tech_LISI", "HarmonyUMAP2D_tech_LISI")
)
#> Error in benchmark_feature_plot(srt = srt, features = features, tool_name = tool_name, reduction = reduction, plot_type = plot_type, plot_boxplot = plot_boxplot, boxplot_jitter = boxplot_jitter, combine = combine, nrow = nrow, ncol = ncol, byrow = byrow, pt.size = pt.size, pt.alpha = pt.alpha, palette = palette, palcolor = palcolor, theme_use = theme_use, theme_args = theme_args, verbose = verbose, ...): The following benchmark columns are missing:
#> "HarmonypcaUMAP2D_tech_LISI" and "HarmonyUMAP2D_tech_LISI"
if (requireNamespace("Signac", quietly = TRUE)) {
data("pbmcmultiome_sub", package = "scop")
pbmcmultiome_sub$batch <- rep(c("batch1", "batch2"), length.out = ncol(pbmcmultiome_sub))
pbmcmultiome_sub <- integration_scop(
pbmcmultiome_sub,
batch = "batch",
assay = "peaks",
integration_method = "Harmony5",
normalization_method = "TFIDF"
)
integration_methods <- c(
"Uncorrected", "Seurat", "CCA", "RPCA", "scVI", "scVI5",
"MNN", "fastMNN", "fastMNN5", "Harmony", "Harmony5",
"Scanorama", "BBKNN", "CSS", "Coralysis", "LIGER", "Conos", "ComBat"
)
p_list <- list()
for (method in integration_methods) {
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = method,
linear_reduction_dims_use = 1:50,
nonlinear_reduction = "umap"
)
p_list[[method]] <- CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = paste0(method, "UMAP2D"),
xlab = "", ylab = "",
title = method,
legend.position = "none",
theme_use = "theme_blank"
)
}
nonlinear_reductions <- c(
"umap", "tsne", "dm", "phate",
"pacmap", "trimap", "largevis", "fr"
)
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "Seurat",
linear_reduction_dims_use = 1:50,
nonlinear_reduction = nonlinear_reductions
)
for (nr in nonlinear_reductions) {
print(
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = paste0("Seurat", nr, "2D"),
xlab = "", ylab = "", title = nr,
legend.position = "none", theme_use = "theme_blank"
)
)
}
}
#> ◌ [2026-05-14 07:55:20] Run integration workflow...
#> ! [2026-05-14 07:55:20] `integration_method = 'Harmony5'` is not compatible with <ChromatinAssay> in current Seurat v5 workflow. Automatically switch to "Harmony"
#> ℹ [2026-05-14 07:55:20] Split `srt_merge` into `srt_list` by "batch"
#> ℹ [2026-05-14 07:55:21] Checking a list of <Seurat>...
#> ! [2026-05-14 07:55:21] Data 1/2 of the `srt_list` is "raw_counts"
#> ℹ [2026-05-14 07:55:21] Perform `RunTFIDF()` on 1/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Perform `FindTopFeatures()` on 1/2 of `srt_list`...
#> ! [2026-05-14 07:55:21] Data 2/2 of the `srt_list` is "raw_counts"
#> ℹ [2026-05-14 07:55:21] Perform `RunTFIDF()` on 2/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Perform `FindTopFeatures()` on 2/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:55:21] Number of available HVF: 2000
#> ℹ [2026-05-14 07:55:21] Finished check
#> ℹ [2026-05-14 07:55:22] `normalization_method` is "TFIDF". Use lsi workflow...
#> ℹ [2026-05-14 07:55:22] Perform linear dimension reduction("svd")
#> Running SVD
#> Scaling cell embeddings
#> ℹ [2026-05-14 07:55:22] Perform Harmony integration
#> ℹ [2026-05-14 07:55:22] Using "Harmonysvd" (2:23) as input
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 't': ‘Z_corr’ is not a valid field or method name for reference class “Rcpp_harmony”
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "LIGER"
)
#> ◌ [2026-05-14 07:54:06] Run integration workflow...
#> ℹ [2026-05-14 07:54:06] Split `srt_merge` into `srt_list` by "tech"
#> ℹ [2026-05-14 07:54:07] Checking a list of <Seurat>...
#> ℹ [2026-05-14 07:54:07] Data 1/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:07] Perform `Seurat::FindVariableFeatures()` on 1/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:08] Data 2/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:08] Perform `Seurat::FindVariableFeatures()` on 2/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:08] Data 3/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:08] Perform `Seurat::FindVariableFeatures()` on 3/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:09] Data 4/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:09] Perform `Seurat::FindVariableFeatures()` on 4/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:09] Data 5/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:09] Perform `Seurat::FindVariableFeatures()` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:10] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:54:10] Number of available HVF: 2000
#> ℹ [2026-05-14 07:54:10] Finished check
#> Warning: Layer ‘ligerScaleData’ is empty
#> ℹ [2026-05-14 07:54:16] Prepare rliger layer "ligerScaleData" ...
#> ℹ [2026-05-14 07:54:17] Perform LIGER integration
#> ℹ [2026-05-14 07:54:23] Adjust neighbor k from 20 to 20 for small-sample clustering
#> ℹ [2026-05-14 07:54:23] Perform `Seurat::FindClusters()` with "louvain"
#> ℹ [2026-05-14 07:54:23] Reorder clusters...
#> ℹ [2026-05-14 07:54:24] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 07:54:24] Perform umap nonlinear dimension reduction using LIGER (1:20)
#> ℹ [2026-05-14 07:54:31] Perform umap nonlinear dimension reduction using LIGER (1:20)
#> ℹ [2026-05-14 07:54:37] Perform umap nonlinear dimension reduction using Uncorrectedpca (1:20)
#> ✔ [2026-05-14 07:54:46] LIGER integration completed
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "Harmony",
compute_lisi = TRUE,
lisi_label_colnames = "tech"
)
#> ◌ [2026-05-14 07:54:46] Run integration workflow...
#> ℹ [2026-05-14 07:54:46] Split `srt_merge` into `srt_list` by "tech"
#> ℹ [2026-05-14 07:54:47] Checking a list of <Seurat>...
#> ℹ [2026-05-14 07:54:48] Data 1/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:48] Perform `Seurat::FindVariableFeatures()` on 1/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:48] Data 2/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:48] Perform `Seurat::FindVariableFeatures()` on 2/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:49] Data 3/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:49] Perform `Seurat::FindVariableFeatures()` on 3/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:49] Data 4/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:49] Perform `Seurat::FindVariableFeatures()` on 4/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:50] Data 5/5 of the `srt_list` has been log-normalized
#> ℹ [2026-05-14 07:54:50] Perform `Seurat::FindVariableFeatures()` on 5/5 of `srt_list`...
#> ℹ [2026-05-14 07:54:50] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:54:51] Number of available HVF: 2000
#> ℹ [2026-05-14 07:54:51] Finished check
#> Warning: Layer ‘scale.data’ is empty
#> ℹ [2026-05-14 07:55:16] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 07:55:16] Perform linear dimension reduction("pca")
#> ℹ [2026-05-14 07:55:17] Perform Harmony integration
#> ℹ [2026-05-14 07:55:17] Using "Harmonypca" (1:20) as input
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 't': ‘Z_corr’ is not a valid field or method name for reference class “Rcpp_harmony”
LISIPlot(
panc8_sub,
features = c("HarmonypcaUMAP2D_tech_LISI", "HarmonyUMAP2D_tech_LISI")
)
#> Error in benchmark_feature_plot(srt = srt, features = features, tool_name = tool_name, reduction = reduction, plot_type = plot_type, plot_boxplot = plot_boxplot, boxplot_jitter = boxplot_jitter, combine = combine, nrow = nrow, ncol = ncol, byrow = byrow, pt.size = pt.size, pt.alpha = pt.alpha, palette = palette, palcolor = palcolor, theme_use = theme_use, theme_args = theme_args, verbose = verbose, ...): The following benchmark columns are missing:
#> "HarmonypcaUMAP2D_tech_LISI" and "HarmonyUMAP2D_tech_LISI"
if (requireNamespace("Signac", quietly = TRUE)) {
data("pbmcmultiome_sub", package = "scop")
pbmcmultiome_sub$batch <- rep(c("batch1", "batch2"), length.out = ncol(pbmcmultiome_sub))
pbmcmultiome_sub <- integration_scop(
pbmcmultiome_sub,
batch = "batch",
assay = "peaks",
integration_method = "Harmony5",
normalization_method = "TFIDF"
)
integration_methods <- c(
"Uncorrected", "Seurat", "CCA", "RPCA", "scVI", "scVI5",
"MNN", "fastMNN", "fastMNN5", "Harmony", "Harmony5",
"Scanorama", "BBKNN", "CSS", "Coralysis", "LIGER", "Conos", "ComBat"
)
p_list <- list()
for (method in integration_methods) {
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = method,
linear_reduction_dims_use = 1:50,
nonlinear_reduction = "umap"
)
p_list[[method]] <- CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = paste0(method, "UMAP2D"),
xlab = "", ylab = "",
title = method,
legend.position = "none",
theme_use = "theme_blank"
)
}
nonlinear_reductions <- c(
"umap", "tsne", "dm", "phate",
"pacmap", "trimap", "largevis", "fr"
)
panc8_sub <- integration_scop(
panc8_sub,
batch = "tech",
integration_method = "Seurat",
linear_reduction_dims_use = 1:50,
nonlinear_reduction = nonlinear_reductions
)
for (nr in nonlinear_reductions) {
print(
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = paste0("Seurat", nr, "2D"),
xlab = "", ylab = "", title = nr,
legend.position = "none", theme_use = "theme_blank"
)
)
}
}
#> ◌ [2026-05-14 07:55:20] Run integration workflow...
#> ! [2026-05-14 07:55:20] `integration_method = 'Harmony5'` is not compatible with <ChromatinAssay> in current Seurat v5 workflow. Automatically switch to "Harmony"
#> ℹ [2026-05-14 07:55:20] Split `srt_merge` into `srt_list` by "batch"
#> ℹ [2026-05-14 07:55:21] Checking a list of <Seurat>...
#> ! [2026-05-14 07:55:21] Data 1/2 of the `srt_list` is "raw_counts"
#> ℹ [2026-05-14 07:55:21] Perform `RunTFIDF()` on 1/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Perform `FindTopFeatures()` on 1/2 of `srt_list`...
#> ! [2026-05-14 07:55:21] Data 2/2 of the `srt_list` is "raw_counts"
#> ℹ [2026-05-14 07:55:21] Perform `RunTFIDF()` on 2/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Perform `FindTopFeatures()` on 2/2 of `srt_list`...
#> ℹ [2026-05-14 07:55:21] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:55:21] Number of available HVF: 2000
#> ℹ [2026-05-14 07:55:21] Finished check
#> ℹ [2026-05-14 07:55:22] `normalization_method` is "TFIDF". Use lsi workflow...
#> ℹ [2026-05-14 07:55:22] Perform linear dimension reduction("svd")
#> Running SVD
#> Scaling cell embeddings
#> ℹ [2026-05-14 07:55:22] Perform Harmony integration
#> ℹ [2026-05-14 07:55:22] Using "Harmonysvd" (2:23) as input
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 't': ‘Z_corr’ is not a valid field or method name for reference class “Rcpp_harmony”