This is a modified version of harmony::RunHarmony specifically designed for compatibility with RunSymphonyMap.
Usage
RunHarmony2(object, ...)
# S3 method for class 'Seurat'
RunHarmony2(
object,
group.by.vars,
assay = NULL,
reduction = "pca",
dims.use = 1:30,
project.dim = TRUE,
reduction.name = "Harmony",
reduction.save = NULL,
reduction.key = "Harmony_",
verbose = TRUE,
seed.use = 11,
...
)Arguments
- object
A Seurat object.
- ...
Additional arguments to be passed to harmony::RunHarmony.
- group.by.vars
The batch variable name.
- assay
Which assay to use. If
NULL, the default assay of the Seurat object will be used. When the object also containsChromatinAssay, the default assay and additionalChromatinAssaywill be preprocessed sequentially.- reduction
Which dimensionality reduction to use. Default is
"pca".- dims.use
The dimensions to be used. Default is
1:30.- project.dim
Whether to project dimension reduction loadings. Default is
TRUE.- reduction.name
The name of the reduction to be stored in the Seurat object. Default is
"Harmony".- reduction.save
Deprecated alias for
reduction.name, retained for compatibility withharmony::RunHarmony.Seurat.- reduction.key
The prefix for the column names of the Harmony embeddings. Default is
"Harmony_".- verbose
Whether to print the message. Default is
TRUE.- seed.use
Random seed for reproducibility. Default is
11.
Examples
data(panc8_sub)
panc8_sub <- standard_scop(panc8_sub)
#> ℹ [2026-05-14 07:27:14] Start standard processing workflow...
#> ℹ [2026-05-14 07:27:14] Checking a list of <Seurat>...
#> ! [2026-05-14 07:27:15] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:27:15] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 07:27:17] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 07:27:17] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:27:17] Number of available HVF: 2000
#> ℹ [2026-05-14 07:27:18] Finished check
#> ℹ [2026-05-14 07:27:18] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 07:27:18] Perform pca linear dimension reduction
#> ℹ [2026-05-14 07:27:19] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 07:27:19] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 07:27:19] Reorder clusters...
#> ℹ [2026-05-14 07:27:19] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 07:27:20] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 07:27:20] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 07:27:26] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 07:27:32] Standard processing workflow completed
panc8_sub <- RunHarmony2(
panc8_sub,
group.by.vars = "tech",
reduction = "pca"
)
#> Transposing data matrix
#> Using automatic lambda estimation
#> Thetas: 2
#> Initializing state using k-means centroids initialization
#> Initializing centroids
#> Harmony 1/10
#> Harmony 2/10
#> Harmony 3/10
#> Harmony 4/10
#> Harmony 5/10
#> Harmony 6/10
#> Harmony 7/10
#> Harmony 8/10
#> Harmony 9/10
#> Harmony converged after 9 iterations
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 't': ‘Z_corr’ is not a valid field or method name for reference class “Rcpp_harmony”
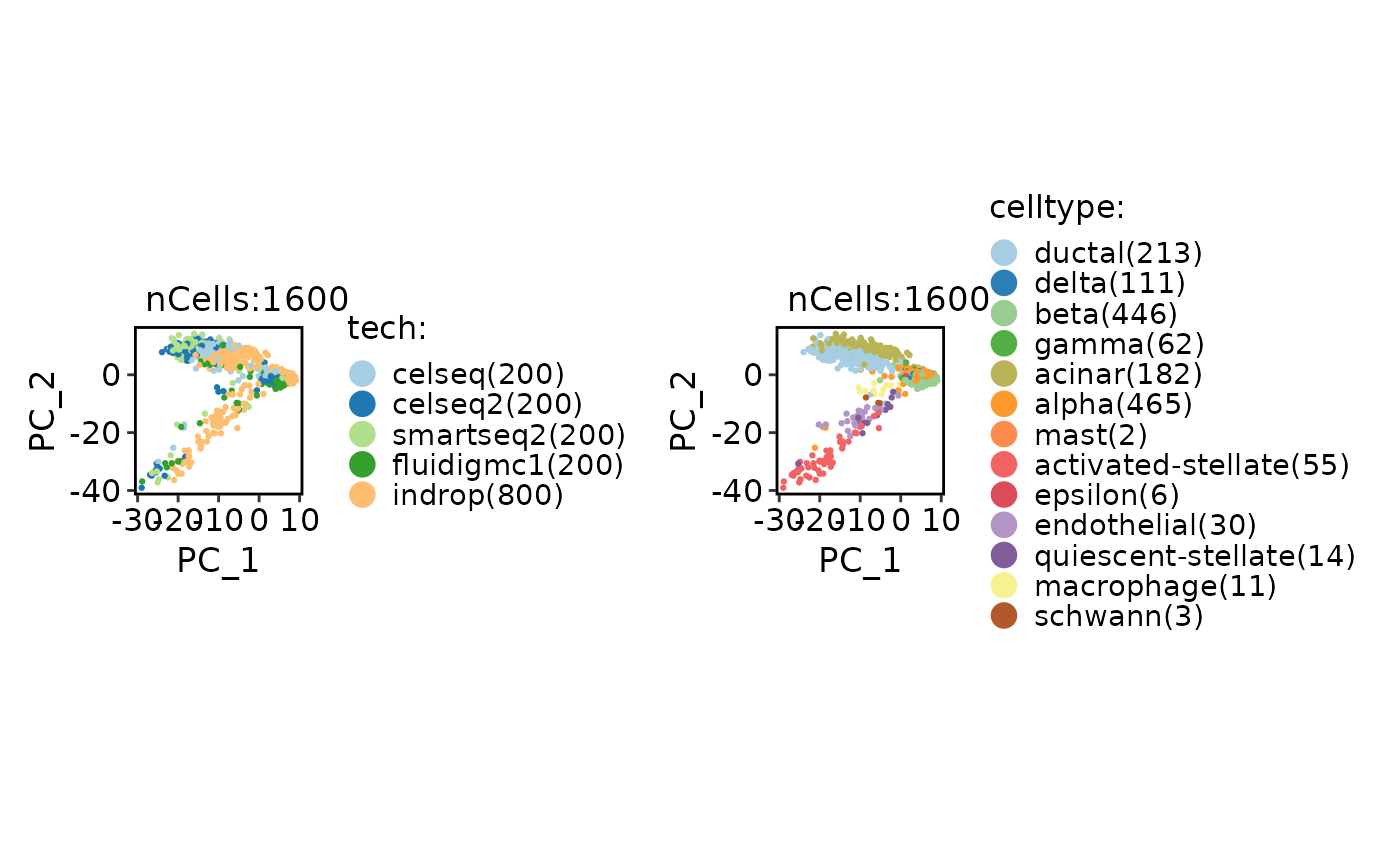
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "pca"
)
 CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "Harmony"
)
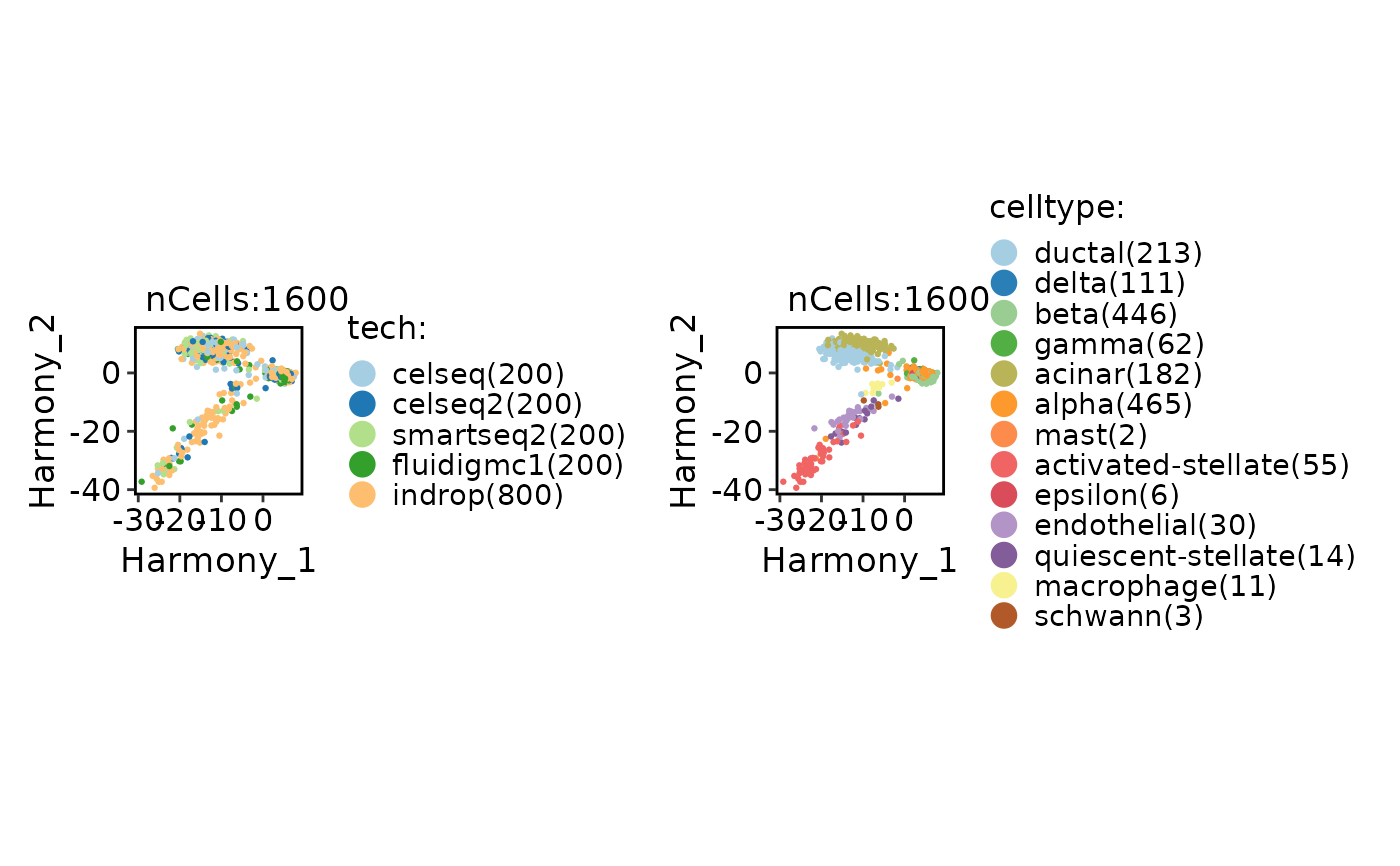
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "Harmony"
)
 panc8_sub <- standard_scop(
panc8_sub,
prefix = "Harmony",
linear_reduction = "Harmony"
)
#> ℹ [2026-05-14 07:27:34] Start standard processing workflow...
#> Error in standard_scop(panc8_sub, prefix = "Harmony", linear_reduction = "Harmony"): `linear_reduction` must be one of: "pca", "svd", "ica", "nmf", "mds",
#> and "glmpca"
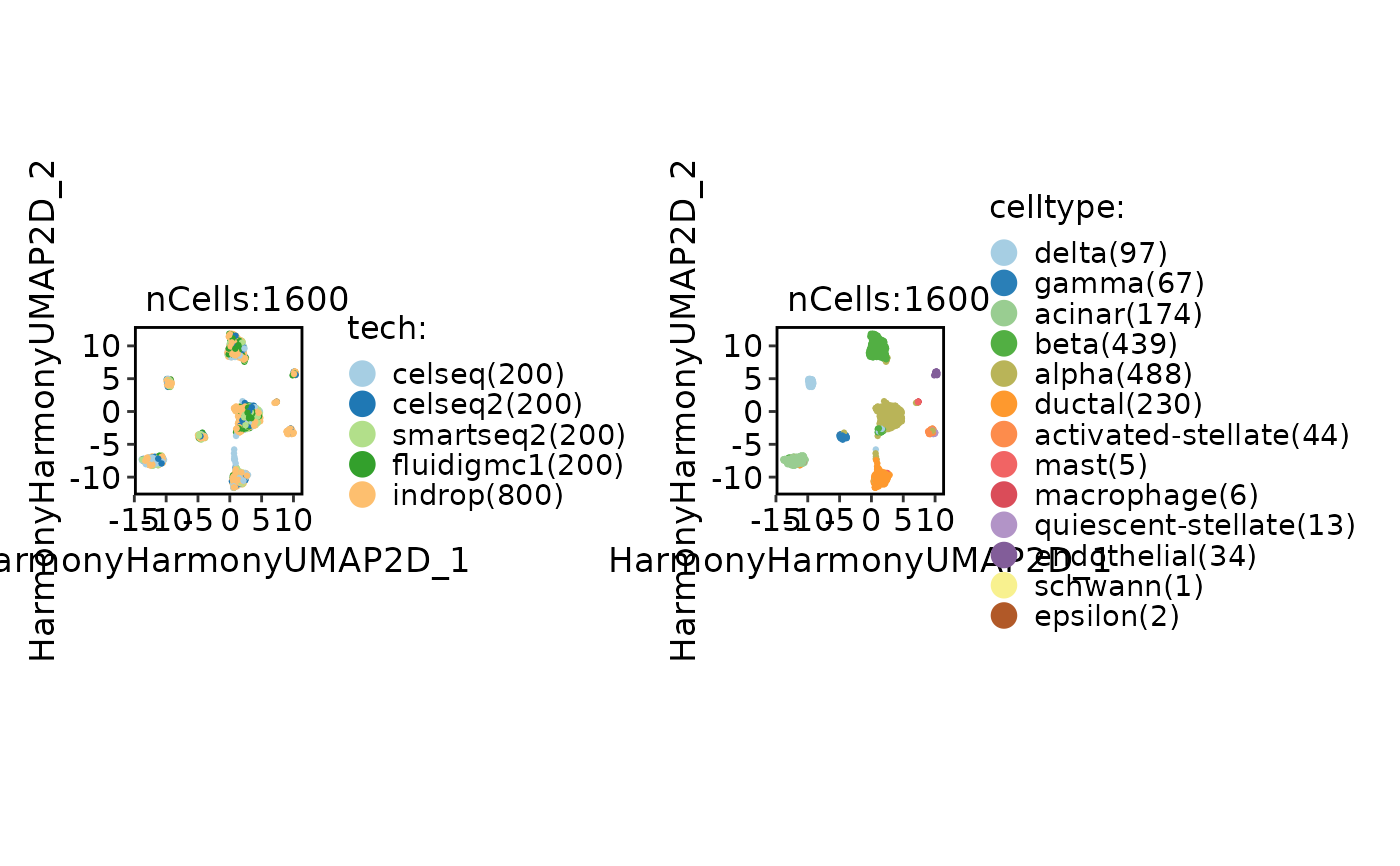
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "StandardpcaUMAP2D"
)
panc8_sub <- standard_scop(
panc8_sub,
prefix = "Harmony",
linear_reduction = "Harmony"
)
#> ℹ [2026-05-14 07:27:34] Start standard processing workflow...
#> Error in standard_scop(panc8_sub, prefix = "Harmony", linear_reduction = "Harmony"): `linear_reduction` must be one of: "pca", "svd", "ica", "nmf", "mds",
#> and "glmpca"
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "StandardpcaUMAP2D"
)
 CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "HarmonyUMAP2D"
)
CellDimPlot(
panc8_sub,
group.by = c("tech", "celltype"),
reduction = "HarmonyUMAP2D"
)