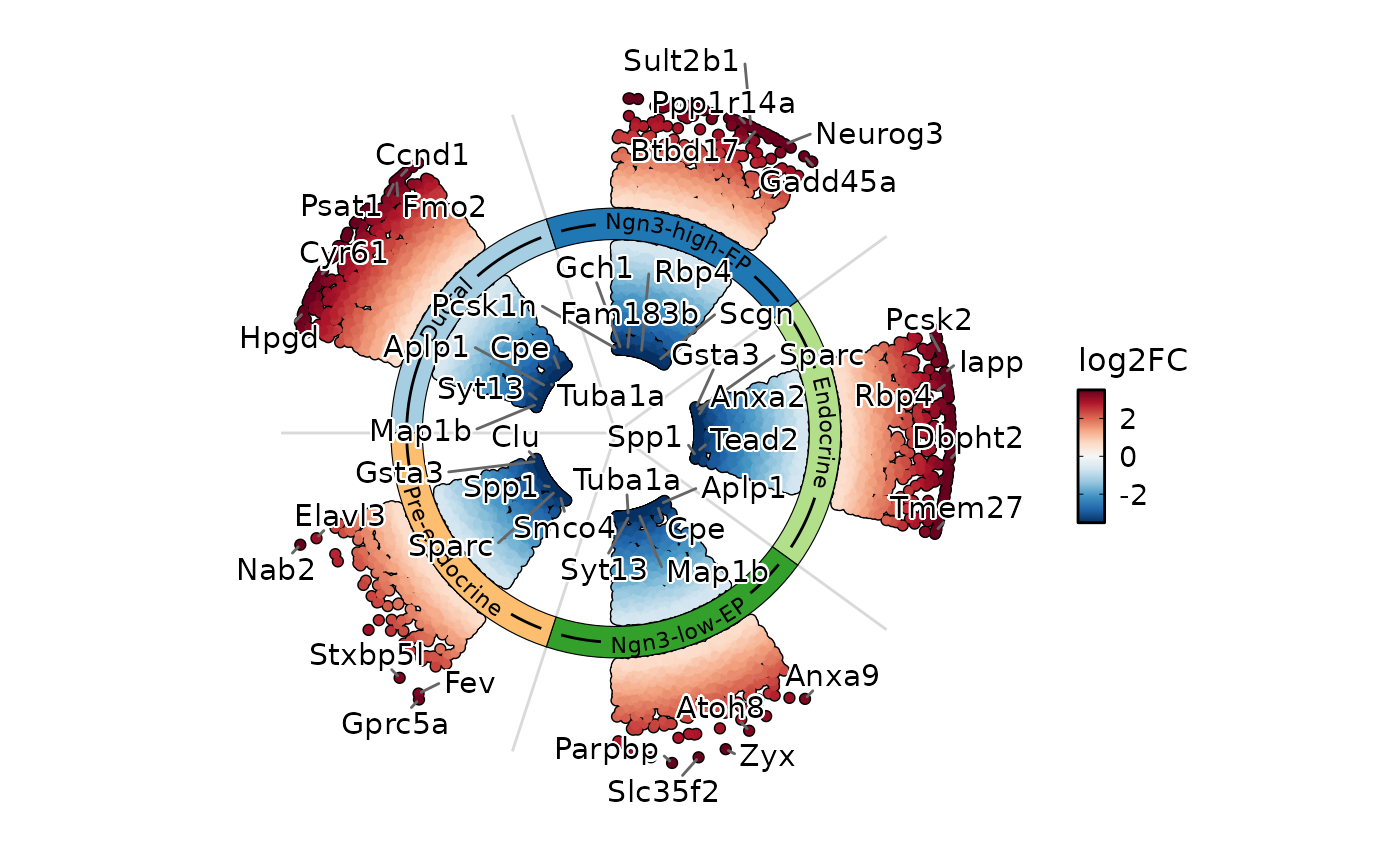
Draw a circular (ring) plot of differential expression results by cell type.
Usage
DEtestRingPlot(
srt,
group.by = NULL,
test.use = "wilcox",
res = NULL,
group_use = NULL,
DE_threshold = "avg_log2FC > 0 & p_val_adj < 0.05",
group_palette = "Chinese",
group_palcolor = NULL,
pt.size = 1,
pt.alpha = 1,
cols.highlight = "black",
sizes.highlight = 1,
alpha.highlight = 1,
stroke.highlight = 0.5,
nlabel = 5,
features_label = NULL,
label.fg = "black",
label.bg = "white",
label.bg.r = 0.1,
label.size = 4,
palette = "RdBu",
palcolor = NULL,
theme_use = "theme_scop",
theme_args = list(),
tile_height = 0.3,
tile_gap = 0.1,
jitter_width = 0.5,
ring_segments = TRUE,
seed = 11
)Arguments
- srt
An object of class
Seuratcontaining the results of differential expression analysis.- group.by
Name of one or more meta.data columns to group (color) cells by.
- test.use
A character string specifying the type of statistical test to use. Default is
"wilcox".- res
A
data.frameordata.tablewith differential expression results. Whenresis provided,srtwill be ignored. The data.frame must contain columns:gene,group1(factor or character),avg_log2FC,p_val_adj, and optionallypct.1andpct.2for calculatingdiff_pct.- group_use
Groups to plot. Default is
NULL(all groups).- DE_threshold
A character string specifying the threshold for differential expression (used to highlight significant genes in all plot types). Default is
"p_val < 0.05"for sample-level methods ("edgeR"and"limma") and"avg_log2FC > 0 & p_val_adj < 0.05"otherwise.- group_palette
Palette for cell types (groups) in Manhattan plot. Default is
"Chinese".- group_palcolor
Custom colors for cell types (groups) in Manhattan plot. Default is
NULL.- pt.size
The size of the points. Default is
1.- pt.alpha
The transparency of the data points. Default is
1.- cols.highlight
A character string specifying the color for highlighted points. Default is
"black".- sizes.highlight
The size of the highlighted points. Default is
1.- alpha.highlight
The transparency of the highlighted points. Default is
1.- stroke.highlight
The stroke width for the highlighted points. Default is
0.5.- nlabel
An integer value specifying the number of labeled points per group. Default is
5.- features_label
A character vector specifying the feature labels to plot. Default is
NULL.- label.fg
A character string specifying the color for the labels' foreground. Default is
"black".- label.bg
A character string specifying the color for the labels' background. Default is
"white".- label.bg.r
The radius of the rounding of the labels' background. Default is
0.1.- label.size
The size of the labels. Default is
4.- palette
Color palette name. Available palettes can be found in thisplot::show_palettes. Default is
"RdBu".- palcolor
Custom colors used to create a color palette. Default is
NULL.- theme_use
Theme to use for the plot. Default is
"theme_scop".- theme_args
A list of additional arguments to pass to the theme function. Default is
list().- tile_height
Height of the cell-type track in ring plot. Default is
0.3.- tile_gap
Gap between the track and nudged points in ring plot. Default is
0.1.- jitter_width
Horizontal jitter range for points in Manhattan plot. Default is
0.5.- ring_segments
Whether to draw segment lines between cell types in ring plot. Default is
TRUE.- seed
Random seed for jitter in ring plot. Default is
11.
Examples
data(pancreas_sub)
pancreas_sub <- standard_scop(pancreas_sub)
#> ℹ [2026-05-14 05:59:29] Start standard processing workflow...
#> ℹ [2026-05-14 05:59:30] Checking a list of <Seurat>...
#> ! [2026-05-14 05:59:30] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 05:59:30] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:59:32] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:59:32] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 05:59:32] Number of available HVF: 2000
#> ℹ [2026-05-14 05:59:32] Finished check
#> ℹ [2026-05-14 05:59:32] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 05:59:33] Perform pca linear dimension reduction
#> ℹ [2026-05-14 05:59:33] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 05:59:34] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 05:59:34] Reorder clusters...
#> ℹ [2026-05-14 05:59:34] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 05:59:34] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 05:59:34] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 05:59:38] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 05:59:42] Standard processing workflow completed
pancreas_sub <- RunDEtest(
pancreas_sub,
group.by = "CellType",
only.pos = FALSE
)
#> ℹ [2026-05-14 05:59:42] Data type is log-normalized
#> ℹ [2026-05-14 05:59:42] Start differential expression test
#> ℹ [2026-05-14 05:59:42] Find all markers(wilcox) among [1] 5 groups...
#> ℹ [2026-05-14 05:59:42] Using 1 core
#> ⠙ [2026-05-14 05:59:42] Running for Ductal [1/5] ■■ 20% | ETA: 1s
#> ✔ [2026-05-14 05:59:42] Completed 5 tasks in 1.2s
#>
#> ℹ [2026-05-14 05:59:42] Building results
#> ✔ [2026-05-14 05:59:44] Differential expression test completed
DEtestRingPlot(
pancreas_sub,
group.by = "CellType"
)