Run Force-Directed Layout (Fruchterman-Reingold algorithm)
Usage
RunFR(object, ...)
# S3 method for class 'Seurat'
RunFR(
object,
reduction = NULL,
dims = NULL,
features = NULL,
assay = NULL,
layer = "data",
graph = NULL,
neighbor = NULL,
k.param = 20,
ndim = 2,
niter = 500,
reduction.name = "FR",
reduction.key = "FR_",
verbose = TRUE,
seed.use = 11L,
...
)
# Default S3 method
RunFR(
object,
assay = NULL,
ndim = 2,
niter = 500,
reduction.key = "FR_",
verbose = TRUE,
seed.use = 11L,
...
)Arguments
- object
An object. This can be a Seurat object, a Neighbor object, or a Graph object. Default is
NULL.- ...
Additional arguments to be passed to igraph::layout_with_fr.
- reduction
Which dimensionality reduction to use. Default is
"pca".- dims
The dimensions to be used. Default is
NULL.- features
A character vector of features to use. Default is
NULL.- assay
Which assay to use. If
NULL, the default assay of the Seurat object will be used. When the object also containsChromatinAssay, the default assay and additionalChromatinAssaywill be preprocessed sequentially.- layer
Which layer to use. Default is
data.- graph
The name of the Graph object to be used. Default is
NULL.- neighbor
The name of the Neighbor object to be used. Default is
NULL.- k.param
The number of nearest neighbors to consider. Default is
20.- ndim
The number of dimensions for the force-directed layout. Default is
2.- niter
The number of iterations for the force-directed layout. Default is
500.- reduction.name
The name of the reduction to be stored in the Seurat object. Default is
"fr".- reduction.key
The prefix for the column names of the force-directed layout embeddings. Default is
"FR_".- verbose
Whether to print the message. Default is
TRUE.- seed.use
Random seed for reproducibility. Default is
11.
Examples
data(pancreas_sub)
pancreas_sub <- standard_scop(pancreas_sub)
#> ℹ [2026-05-14 07:21:59] Start standard processing workflow...
#> ℹ [2026-05-14 07:22:00] Checking a list of <Seurat>...
#> ! [2026-05-14 07:22:00] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 07:22:00] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 07:22:02] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 07:22:03] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 07:22:03] Number of available HVF: 2000
#> ℹ [2026-05-14 07:22:03] Finished check
#> ℹ [2026-05-14 07:22:03] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 07:22:03] Perform pca linear dimension reduction
#> ℹ [2026-05-14 07:22:04] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 07:22:04] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 07:22:04] Reorder clusters...
#> ℹ [2026-05-14 07:22:05] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 07:22:05] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 07:22:05] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 07:22:10] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 07:22:16] Standard processing workflow completed
pancreas_sub <- RunFR(
object = pancreas_sub,
features = SeuratObject::VariableFeatures(pancreas_sub)
)
#> ℹ [2026-05-14 07:22:16] Running force-directed layout
#> ℹ [2026-05-14 07:22:16] Computing nearest neighbor graph and SNN
#> ℹ [2026-05-14 07:22:29] Force-directed layout computed
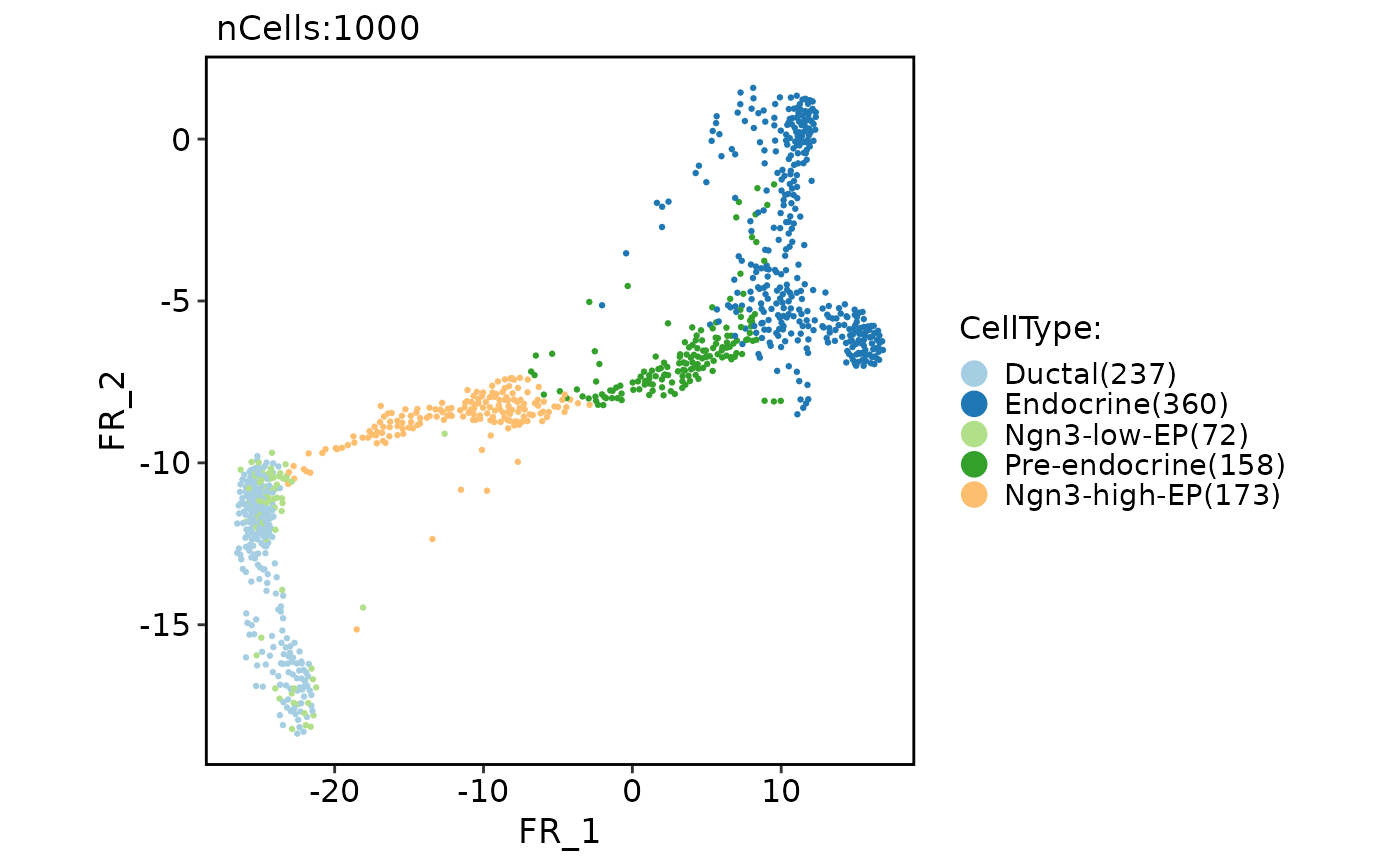
CellDimPlot(
pancreas_sub,
group.by = "CellType",
reduction = "fr"
)