CCC statistical distribution and summary plots
Usage
CCCStatPlot(
srt,
method = NULL,
condition = NULL,
dataset = 1,
comparison = c(1, 2),
plot_type = c("bar", "sankey", "box", "violin", "role_scatter", "role_network",
"role_network_marsilea", "pathway_summary", "comparison", "lr_contribution", "gene",
"ranknet", "scatter", "role_change"),
display_by = c("aggregation", "interaction"),
sender.use = NULL,
receiver.use = NULL,
ligand.use = NULL,
receptor.use = NULL,
interaction.use = NULL,
signaling = NULL,
pairLR.use = NULL,
slot.name = "net",
thresh = 0.05,
measure = c("count", "weight"),
pattern = c("all", "outgoing", "incoming"),
compare_by = c("overall", "celltype"),
value = "score",
stat_type = c("score", "count"),
top_n = 20,
x_text_angle = 90,
min_receiver_flow = 0,
link_alpha = 0.6,
facet_by = NULL,
edge_value = c("sum", "mean", "max", "count"),
edge_threshold = 0,
palette = "Chinese",
palcolor = NULL,
cell_palette = NULL,
cell_palcolor = NULL,
link_palette = NULL,
link_palcolor = NULL,
title = NULL,
subtitle = NULL,
legend.position = "right",
legend.direction = "vertical",
font.size = 10,
theme_use = "theme_scop",
theme_args = list(),
grid_major = TRUE,
grid_major_colour = "grey80",
grid_major_linetype = 2,
grid_major_linewidth = 0.3,
combine = TRUE,
nrow = NULL,
ncol = NULL,
verbose = TRUE,
...
)Arguments
- srt
A
Seuratobject.- method
Communication result type to use.
- condition
Result name or comparison name.
- dataset
Dataset index or name.
- comparison
Comparison indices or names.
- plot_type
Plot type. One of:
"bar"— horizontal bar chart of top pairs or interactions according todisplay_by."sankey"— alluvial/sankey flow diagram."box"/"violin"— distribution of interaction scores across sender-receiver pairs."comparison"— comparison bars at overall or celltype level."lr_contribution"— ligand-receptor contribution bar plot."gene"— pathway-related ligand/receptor gene expression panel."ranknet"— pathway ranking comparison plot."scatter"— outgoing vs. incoming signaling strength scatter."role_change"— signaling change scatter for one cell identity.
- display_by
Whether to summarize by
"aggregation"or"interaction".- sender.use
Sender cell types to keep.
- receiver.use
Receiver cell types to keep.
- ligand.use
Ligands to keep.
- receptor.use
Receptors to keep.
- interaction.use
Interaction names to keep.
- signaling
Signaling pathway to focus on.
- pairLR.use
Specific ligand-receptor pair(s) to keep.
- slot.name
CellChat slot name.
- thresh
Significance threshold used when extracting communication results.
- measure
Summary measure for CellChat objects.
- pattern
Pattern used for pathway role plots.
- compare_by
Comparison mode for CellChat summary plots.
- value
Value column or summary statistic to use.
- stat_type
For
"bar": what to summarize per interaction. One of"score"(total aggregated score) or"count"(number of significant interactions).- top_n
Number of top records to retain.
- x_text_angle
Rotation angle for x-axis labels.
- min_receiver_flow
For
"sankey": minimum total receiver-side flow retained after top-N ranking. Useful when many small receiver nodes make the right side unreadable.- link_alpha
Alpha used for network edges.
- facet_by
Faceting variable for interaction-level plots.
- edge_value
Aggregation statistic for network edges.
- edge_threshold
Minimum edge value to keep.
- palette
Main palette name.
- palcolor
Main custom palette colors.
- cell_palette
Cell annotation palette name.
- cell_palcolor
Custom cell annotation colors.
- link_palette
Link palette name.
- link_palcolor
Custom link palette colors.
- title
Plot title.
- subtitle
Plot subtitle.
- legend.position
Legend position.
- legend.direction
Legend direction.
- font.size
Base font size.
- theme_use
Theme function used for styling.
- theme_args
Arguments passed to the theme function.
- grid_major
Whether to show major panel grid lines for applicable statistical panels. Default is
TRUE.- grid_major_colour
Color of major panel grid lines.
- grid_major_linetype
Linetype of major panel grid lines.
- grid_major_linewidth
Line width of major panel grid lines.
- combine
Whether to combine multiple panels.
- nrow
Number of rows in combined layout.
- ncol
Number of columns in combined layout.
- verbose
Whether to print messages.
- ...
Additional plot-specific options.
Examples
if (requireNamespace("CellChat", quietly = TRUE)) {
data(pancreas_sub)
pancreas_sub <- standard_scop(pancreas_sub)
pc1 <- Seurat::Embeddings(pancreas_sub, "Standardpca")[, 1]
ct <- as.character(pancreas_sub$CellType)
ct_medians <- tapply(pc1, ct, median)
pancreas_sub$Condition <- ifelse(
pc1 > ct_medians[ct],
"ConditionA",
"ConditionB"
)
pancreas_sub <- RunCellChat(
pancreas_sub,
group.by = "CellType",
group_column = "Condition",
group_cmp = list(c("ConditionA", "ConditionB")),
species = "Mus_musculus"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "sankey",
display_by = "aggregation",
top_n = 20
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "sankey",
display_by = "interaction",
top_n = 20
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "box",
facet_by = "sender",
top_n = 200
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "violin",
facet_by = "receiver",
top_n = 200
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "bar",
palette = "Paired",
top_n = 100
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "scatter"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "lr_contribution",
signaling = "MK"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "gene",
signaling = "MK"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "comparison",
measure = "count",
compare_by = "overall"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "comparison",
measure = "weight",
compare_by = "celltype",
pattern = "all"
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "ranknet"
)
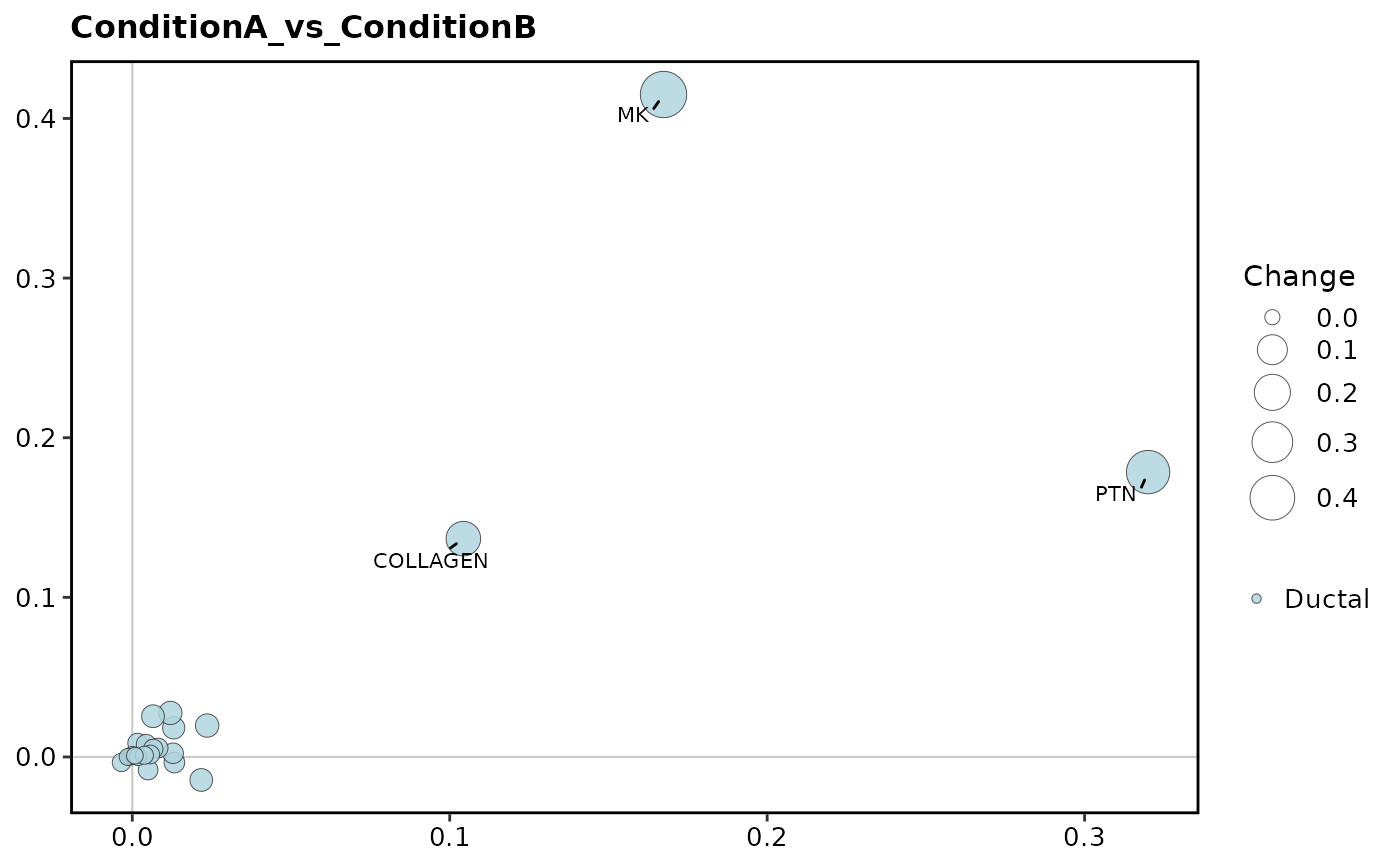
CCCStatPlot(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
idents.use = "Ductal",
plot_type = "role_change"
)
}
#> ℹ [2026-05-14 05:32:43] Start standard processing workflow...
#> ℹ [2026-05-14 05:32:44] Checking a list of <Seurat>...
#> ! [2026-05-14 05:32:44] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 05:32:44] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:32:45] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:32:45] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 05:32:45] Number of available HVF: 2000
#> ℹ [2026-05-14 05:32:46] Finished check
#> ℹ [2026-05-14 05:32:46] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 05:32:46] Perform pca linear dimension reduction
#> ℹ [2026-05-14 05:32:46] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 05:32:47] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 05:32:47] Reorder clusters...
#> ℹ [2026-05-14 05:32:47] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 05:32:47] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 05:32:47] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 05:32:50] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 05:32:53] Standard processing workflow completed
#> ℹ [2026-05-14 05:32:53] Start CellChat analysis
#> ℹ [2026-05-14 05:32:53] Processing condition: "ConditionA"
#> [1] "Create a CellChat object from a data matrix"
#> Set cell identities for the new CellChat object
#> The cell groups used for CellChat analysis are Ductal, Ngn3-high-EP, Endocrine, Ngn3-low-EP, Pre-endocrine
#> The number of highly variable ligand-receptor pairs used for signaling inference is 542
#> triMean is used for calculating the average gene expression per cell group.
#> [1] ">>> Run CellChat on sc/snRNA-seq data <<< [2026-05-14 05:32:54.259776]"
#> [1] ">>> CellChat inference is done. Parameter values are stored in `object@options$parameter` <<< [2026-05-14 05:33:12.642106]"
#> ℹ [2026-05-14 05:33:12] Processing condition: "ConditionB"
#> [1] "Create a CellChat object from a data matrix"
#> Set cell identities for the new CellChat object
#> The cell groups used for CellChat analysis are Endocrine, Ngn3-high-EP, Ductal, Ngn3-low-EP, Pre-endocrine
#> The number of highly variable ligand-receptor pairs used for signaling inference is 601
#> triMean is used for calculating the average gene expression per cell group.
#> [1] ">>> Run CellChat on sc/snRNA-seq data <<< [2026-05-14 05:33:13.768842]"
#> [1] ">>> CellChat inference is done. Parameter values are stored in `object@options$parameter` <<< [2026-05-14 05:33:33.039533]"
#> ℹ [2026-05-14 05:33:33] Merging CellChat objects for comparison "ConditionA_vs_ConditionB"
#> Merge the following slots: 'data.signaling','images','net', 'netP','meta', 'idents', 'var.features' , 'DB', and 'LR'.
#> ✔ [2026-05-14 05:33:33] CellChat analysis completed
#> ! [2026-05-14 05:33:41] `thisplot::StatPlot()` sankey is count-based. For `CCCStatPlot()` with `plot_type = 'sankey'`, `edge_value` is used to rank/filter pairs, but flow width is shown by interaction count.
#> ℹ [2026-05-14 05:33:44] Setting `group.by` to "Features" as `plot.by` is set to "feature"