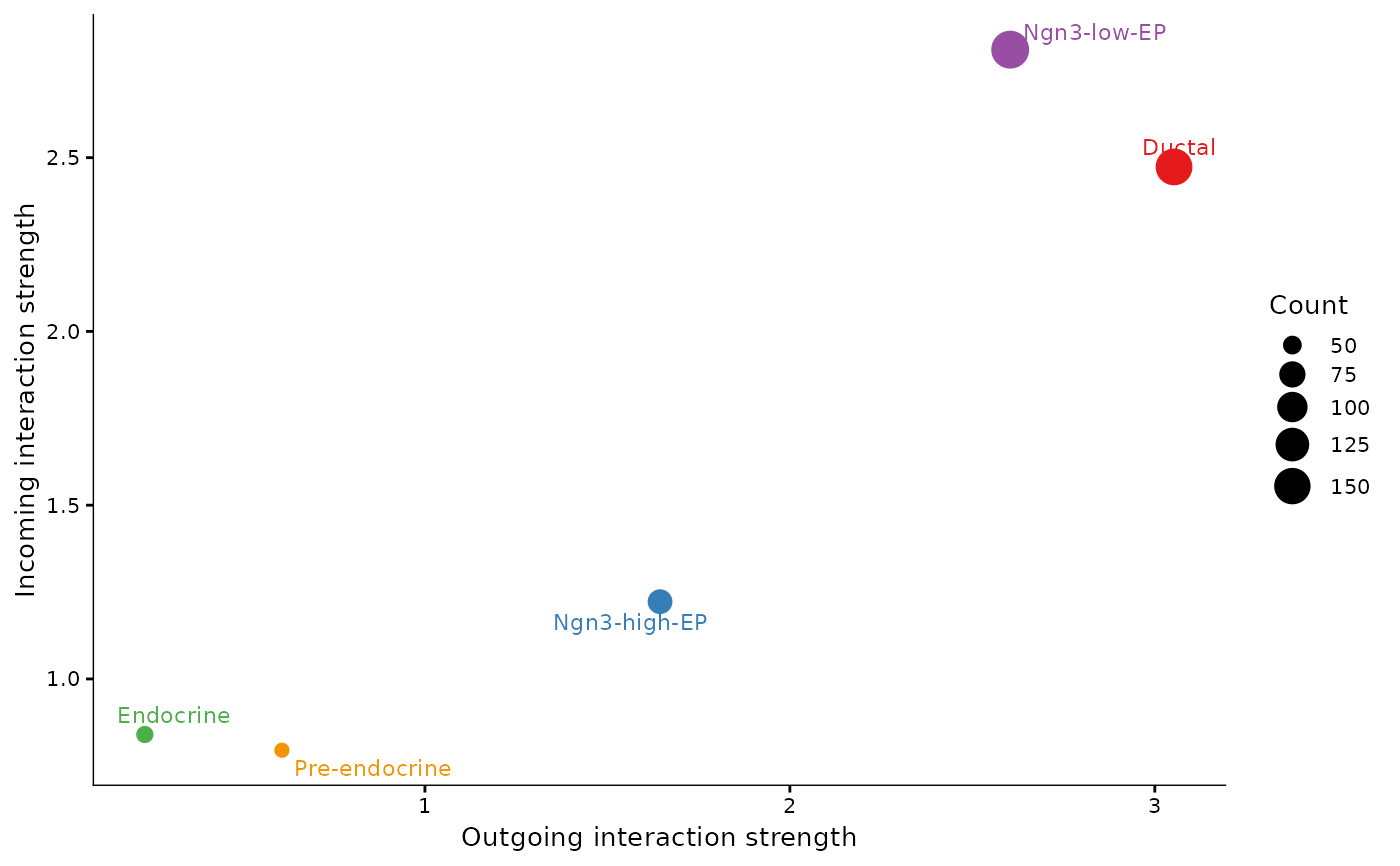
Run CellChat analysis
Usage
RunCellChat(
srt,
group.by,
species = c("Homo_sapiens", "Mus_musculus", "zebrafish"),
split.by = NULL,
annotation_selected = NULL,
group_column = NULL,
group_cmp = NULL,
thresh = 0.05,
min.cells = 10,
assay = NULL,
layer = "data",
verbose = TRUE
)Arguments
- srt
A Seurat object.
- group.by
Name of one or more meta.data columns to group (color) cells by.
- species
The species of the data, either
"Homo_sapiens","Mus_musculus", or"zebrafish".- split.by
Name of a column in meta.data column to split plot by. Default is
NULL.- annotation_selected
A vector of cell annotations of interest for running the
CellChatanalysis. If not provided, all cell types will be considered.- group_column
Name of the metadata column in the
Seuratobject that defines conditions or groups.- group_cmp
A list of pairwise condition comparisons for differential
CellChatanalysis.- thresh
The threshold for computing centrality scores. Default is
0.05.- min.cells
the minmum number of expressed cells required for the genes that are considered for cell-cell communication analysis. Default is
10.- assay
Which assay to use. If
NULL, the default assay of theSeuratobject will be used.- layer
The layer to use for the expression data. Default is
"data".- verbose
Whether to print the message. Default is
TRUE.
Examples
data(pancreas_sub)
pancreas_sub <- standard_scop(pancreas_sub)
#> ℹ [2026-05-14 06:55:00] Start standard processing workflow...
#> ℹ [2026-05-14 06:55:01] Checking a list of <Seurat>...
#> ! [2026-05-14 06:55:01] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 06:55:01] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 06:55:03] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 06:55:03] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 06:55:04] Number of available HVF: 2000
#> ℹ [2026-05-14 06:55:04] Finished check
#> ℹ [2026-05-14 06:55:04] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 06:55:04] Perform pca linear dimension reduction
#> ℹ [2026-05-14 06:55:04] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 06:55:05] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 06:55:05] Reorder clusters...
#> ℹ [2026-05-14 06:55:05] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 06:55:05] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 06:55:05] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 06:55:10] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 06:55:15] Standard processing workflow completed
pancreas_sub <- RunCellChat(
pancreas_sub,
group.by = "CellType",
species = "Mus_musculus"
)
#> ℹ [2026-05-14 06:55:15] Start CellChat analysis
#> [1] "Create a CellChat object from a data matrix"
#> Set cell identities for the new CellChat object
#> The cell groups used for CellChat analysis are Ductal, Ngn3-high-EP, Endocrine, Ngn3-low-EP, Pre-endocrine
#> The number of highly variable ligand-receptor pairs used for signaling inference is 841
#> triMean is used for calculating the average gene expression per cell group.
#> [1] ">>> Run CellChat on sc/snRNA-seq data <<< [2026-05-14 06:55:16.649211]"
#> [1] ">>> CellChat inference is done. Parameter values are stored in `object@options$parameter` <<< [2026-05-14 06:55:38.676401]"
#> ✔ [2026-05-14 06:55:38] CellChat analysis completed
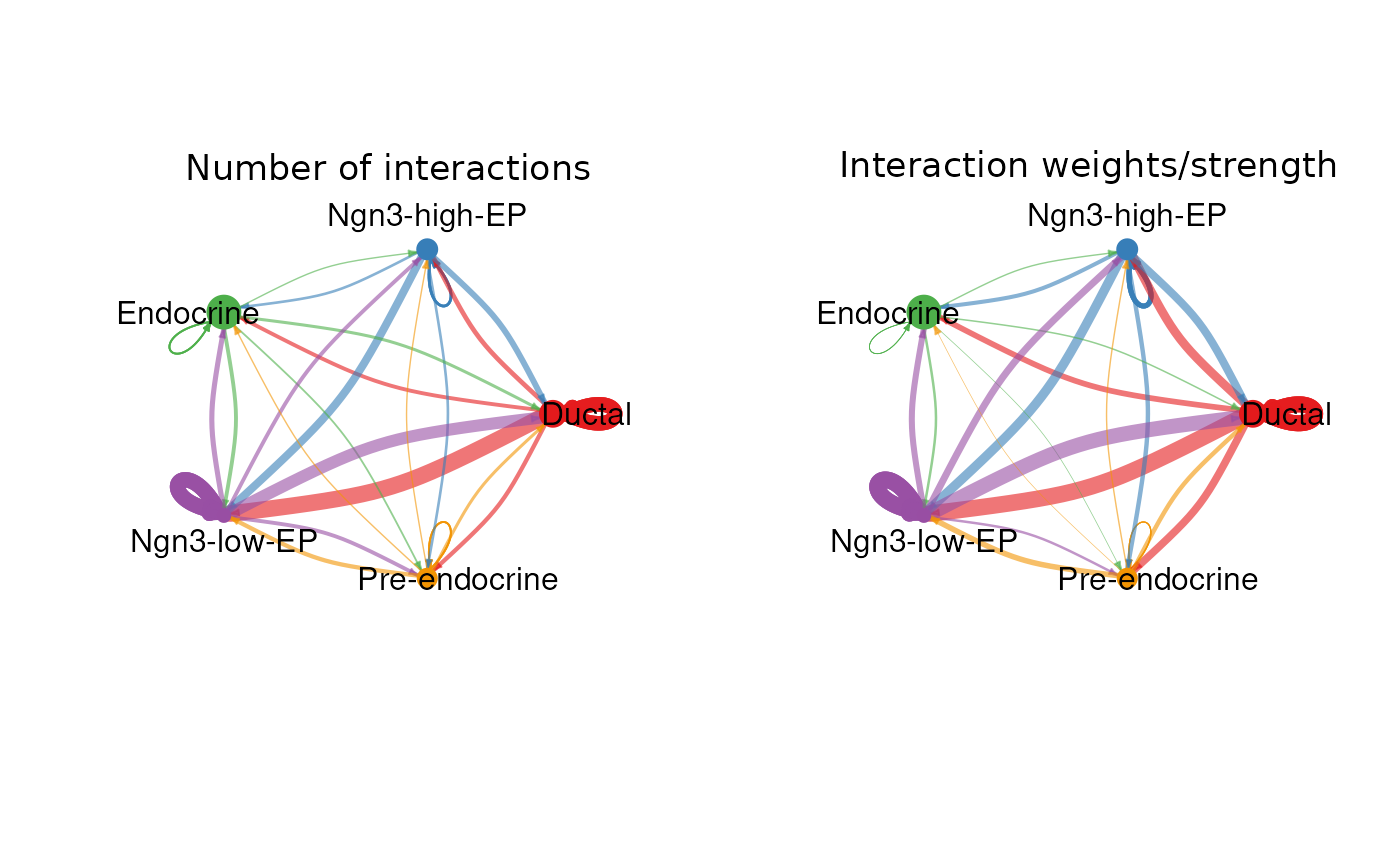
CCCNetworkPlot(
pancreas_sub,
method = "CellChat",
plot_type = "bipartite"
)
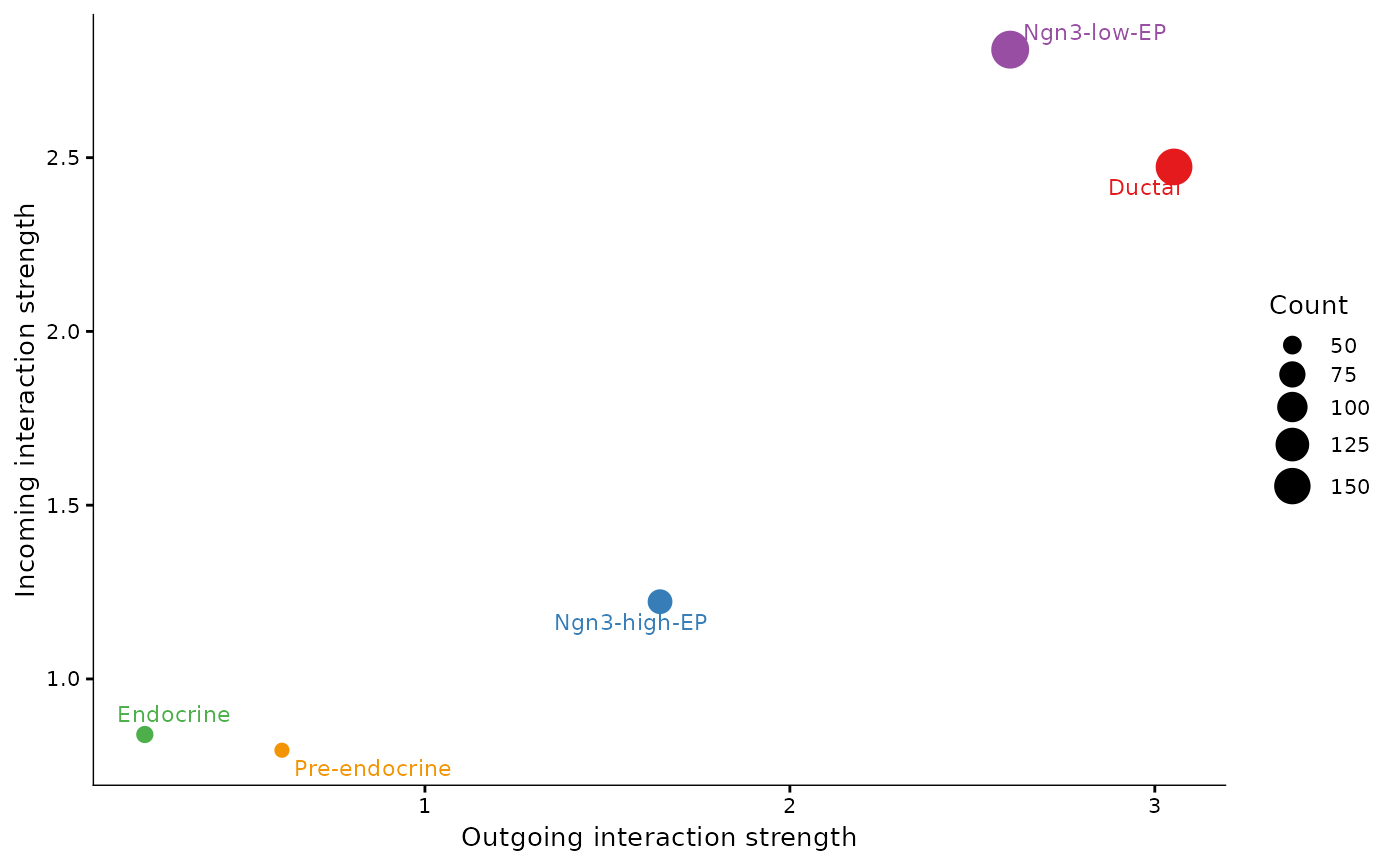
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
plot_type = "heatmap"
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
plot_type = "heatmap"
)
 CCCStatPlot(
pancreas_sub,
method = "CellChat",
plot_type = "violin",
top_n = 50
)
CCCStatPlot(
pancreas_sub,
method = "CellChat",
plot_type = "violin",
top_n = 50
)