CCC heatmap and dot matrix plot
Usage
CCCHeatmap(
srt,
method = NULL,
condition = NULL,
dataset = 1,
comparison = c(1, 2),
plot_type = c("heatmap", "focused_heatmap", "dot", "matrix_dot", "tile",
"source_target_dot", "source_target_tile", "sample_dot", "bubble", "bubble_lr",
"pathway_bubble", "ligand_target", "role_heatmap", "role_network",
"role_network_marsilea", "diff_heatmap"),
display_by = c("aggregation", "interaction"),
sender.use = NULL,
receiver.use = NULL,
ligand.use = NULL,
receptor.use = NULL,
interaction.use = NULL,
signaling = NULL,
pairLR.use = NULL,
slot.name = "net",
thresh = 0.05,
measure = c("count", "weight"),
pattern = c("outgoing", "incoming", "all"),
value = "sum",
top_n = 500,
top_anno = "bar",
right_anno = "cell",
left_anno = "bar",
bottom_anno = "cell",
bar_value = "sum",
add_text = NULL,
cluster_rows = FALSE,
cluster_columns = FALSE,
color.by = c("score", "pvalue"),
x_text_angle = 90,
facet_by = NULL,
show_row_names = TRUE,
show_column_names = TRUE,
edge_value = c("sum", "mean", "max", "count"),
border = TRUE,
value_palette = "RdBu",
value_palcolor = NULL,
cell_palette = "Chinese",
cell_palcolor = NULL,
palette = "Chinese",
palcolor = NULL,
width = NULL,
height = NULL,
units = "inch",
title = NULL,
subtitle = NULL,
legend.position = "right",
legend.direction = "vertical",
font.size = 10,
theme_use = "theme_scop",
theme_args = list(),
verbose = TRUE,
...
)Arguments
- srt
A
Seuratobject.- method
Communication result type to use.
- condition
Result name or comparison name.
- dataset
Dataset index or name.
- comparison
Comparison indices or names.
- plot_type
Plot type. One of
"heatmap"or"dot"."bubble"is a CellChat-specific interaction bubble matrix."ligand_target"is a special heatmap path available only withNicheNet/MultiNicheNetresults."role_heatmap"and"diff_heatmap"are CellChat-specific pathway role views.- display_by
Whether to summarize by
"aggregation"or"interaction".- sender.use
Sender cell types to keep.
- receiver.use
Receiver cell types to keep.
- ligand.use
Ligands to keep.
- receptor.use
Receptors to keep.
- interaction.use
Interaction names to keep.
- signaling
Signaling pathway to focus on.
- pairLR.use
Specific ligand-receptor pair(s) to keep.
- slot.name
CellChat slot name.
- thresh
Significance threshold used when extracting communication results.
- measure
Summary measure for CellChat objects.
- pattern
Pattern used for pathway role plots.
- value
Value column or summary statistic to use.
- top_n
Number of top records to retain.
- top_anno, bottom_anno
Column-side annotations for sender groups. Each side accepts
NULL,"bar","box","point","line","histogram","density","violin","cell", or a character vector containing multiple values. Defaults aretop_anno = "bar"andbottom_anno = "cell".- left_anno, right_anno
Row-side annotations for receiver groups. Each side accepts
NULL,"bar","box","point","line","histogram","density","violin","cell", or a character vector containing multiple values. Defaults areleft_anno = "bar"andright_anno = "cell".- bar_value
Aggregation metric shown in the bar annotations. One or more of
"count","sum","mean", or"max". Multiple values add multiple annotation tracks. Default"sum".- add_text
Logical. Show numeric value labels inside each cell (heatmap mode only). Default
TRUEfor aggregation mode,FALSEfor interaction mode.- cluster_rows, cluster_columns
Whether to cluster heatmap rows/columns. Defaults are both
FALSE.- color.by
For interaction heatmaps, value used for tile coloring. Usually
"score"or"pvalue".- x_text_angle
Rotation angle for x-axis labels.
- facet_by
Faceting variable for interaction-level plots.
- show_row_names, show_column_names
Whether to draw row/column names for the heatmap body.
- edge_value
Aggregation statistic for network edges.
- border
Logical. Whether to draw borders for the heatmap body and all annotation tracks. Default
TRUE.- value_palette
Palette used for heatmap value fills.
- value_palcolor
Optional custom colors for
value_palette.- cell_palette
Cell annotation palette name.
- cell_palcolor
Custom cell annotation colors.
- palette
Main palette name.
- palcolor
Main custom palette colors.
- width, height
Optional heatmap body width and height. When only one is supplied, the other is inferred from the matrix dimensions to keep cells square. When both are
NULL, a square-cell size is computed automatically.- units
Units for
widthandheight. Default"inch".- title
Plot title.
- subtitle
Plot subtitle.
- legend.position
Legend position.
- legend.direction
Legend direction.
- font.size
Base font size.
- theme_use
Theme function used for styling.
- theme_args
Arguments passed to the theme function.
- verbose
Whether to print messages.
- ...
Additional plot-specific options.
Examples
data(pancreas_sub)
pancreas_sub <- standard_scop(pancreas_sub)
#> ℹ [2026-05-14 05:24:02] Start standard processing workflow...
#> ℹ [2026-05-14 05:24:04] Checking a list of <Seurat>...
#> ! [2026-05-14 05:24:04] Data 1/1 of the `srt_list` is "unknown"
#> ℹ [2026-05-14 05:24:04] Perform `NormalizeData()` with `normalization.method = 'LogNormalize'` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:24:05] Perform `Seurat::FindVariableFeatures()` on 1/1 of `srt_list`...
#> ℹ [2026-05-14 05:24:05] Use the separate HVF from `srt_list`
#> ℹ [2026-05-14 05:24:05] Number of available HVF: 2000
#> ℹ [2026-05-14 05:24:06] Finished check
#> ℹ [2026-05-14 05:24:06] Perform `Seurat::ScaleData()`
#> ℹ [2026-05-14 05:24:06] Perform pca linear dimension reduction
#> ℹ [2026-05-14 05:24:16] Use stored estimated dimensions 1:20 for Standardpca
#> ℹ [2026-05-14 05:24:17] Perform `Seurat::FindClusters()` with `cluster_algorithm = 'louvain'` and `cluster_resolution = 0.6`
#> ℹ [2026-05-14 05:24:17] Reorder clusters...
#> ℹ [2026-05-14 05:24:17] Skip `log1p()` because `layer = data` is not "counts"
#> ℹ [2026-05-14 05:24:17] Perform umap nonlinear dimension reduction
#> ℹ [2026-05-14 05:24:17] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ℹ [2026-05-14 05:24:20] Perform umap nonlinear dimension reduction using Standardpca (1:20)
#> ✔ [2026-05-14 05:24:23] Standard processing workflow completed
pc1 <- Seurat::Embeddings(pancreas_sub, "Standardpca")[, 1]
ct <- as.character(pancreas_sub$CellType)
ct_medians <- tapply(pc1, ct, median)
pancreas_sub$Condition <- ifelse(
pc1 > ct_medians[ct],
"ConditionA",
"ConditionB"
)
pancreas_sub <- RunCellChat(
pancreas_sub,
group.by = "CellType",
group_column = "Condition",
group_cmp = list(c("ConditionA", "ConditionB")),
species = "Mus_musculus"
)
#> ℹ [2026-05-14 05:24:23] Start CellChat analysis
#> ℹ [2026-05-14 05:27:21] Processing condition: "ConditionA"
#> [1] "Create a CellChat object from a data matrix"
#> Set cell identities for the new CellChat object
#> The cell groups used for CellChat analysis are Ductal, Ngn3-high-EP, Endocrine, Ngn3-low-EP, Pre-endocrine
#> The number of highly variable ligand-receptor pairs used for signaling inference is 542
#> triMean is used for calculating the average gene expression per cell group.
#> [1] ">>> Run CellChat on sc/snRNA-seq data <<< [2026-05-14 05:27:22.885775]"
#> [1] ">>> CellChat inference is done. Parameter values are stored in `object@options$parameter` <<< [2026-05-14 05:27:41.813448]"
#> ℹ [2026-05-14 05:27:41] Processing condition: "ConditionB"
#> [1] "Create a CellChat object from a data matrix"
#> Set cell identities for the new CellChat object
#> The cell groups used for CellChat analysis are Endocrine, Ngn3-high-EP, Ductal, Ngn3-low-EP, Pre-endocrine
#> The number of highly variable ligand-receptor pairs used for signaling inference is 601
#> triMean is used for calculating the average gene expression per cell group.
#> [1] ">>> Run CellChat on sc/snRNA-seq data <<< [2026-05-14 05:27:43.049499]"
#> [1] ">>> CellChat inference is done. Parameter values are stored in `object@options$parameter` <<< [2026-05-14 05:28:03.373663]"
#> ℹ [2026-05-14 05:28:03] Merging CellChat objects for comparison "ConditionA_vs_ConditionB"
#> Merge the following slots: 'data.signaling','images','net', 'netP','meta', 'idents', 'var.features' , 'DB', and 'LR'.
#> ✔ [2026-05-14 05:28:03] CellChat analysis completed
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "aggregation",
top_n = 20
)
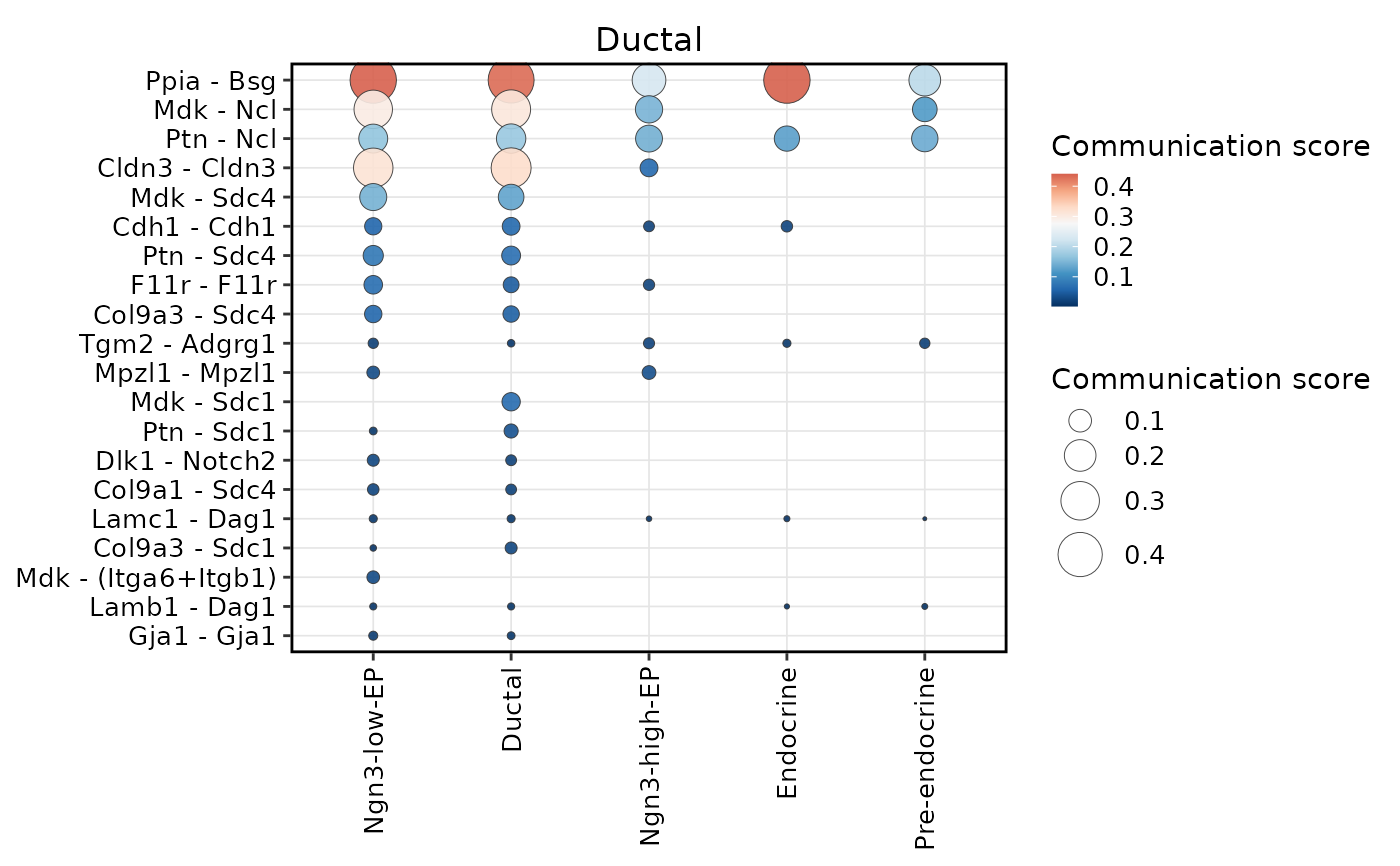
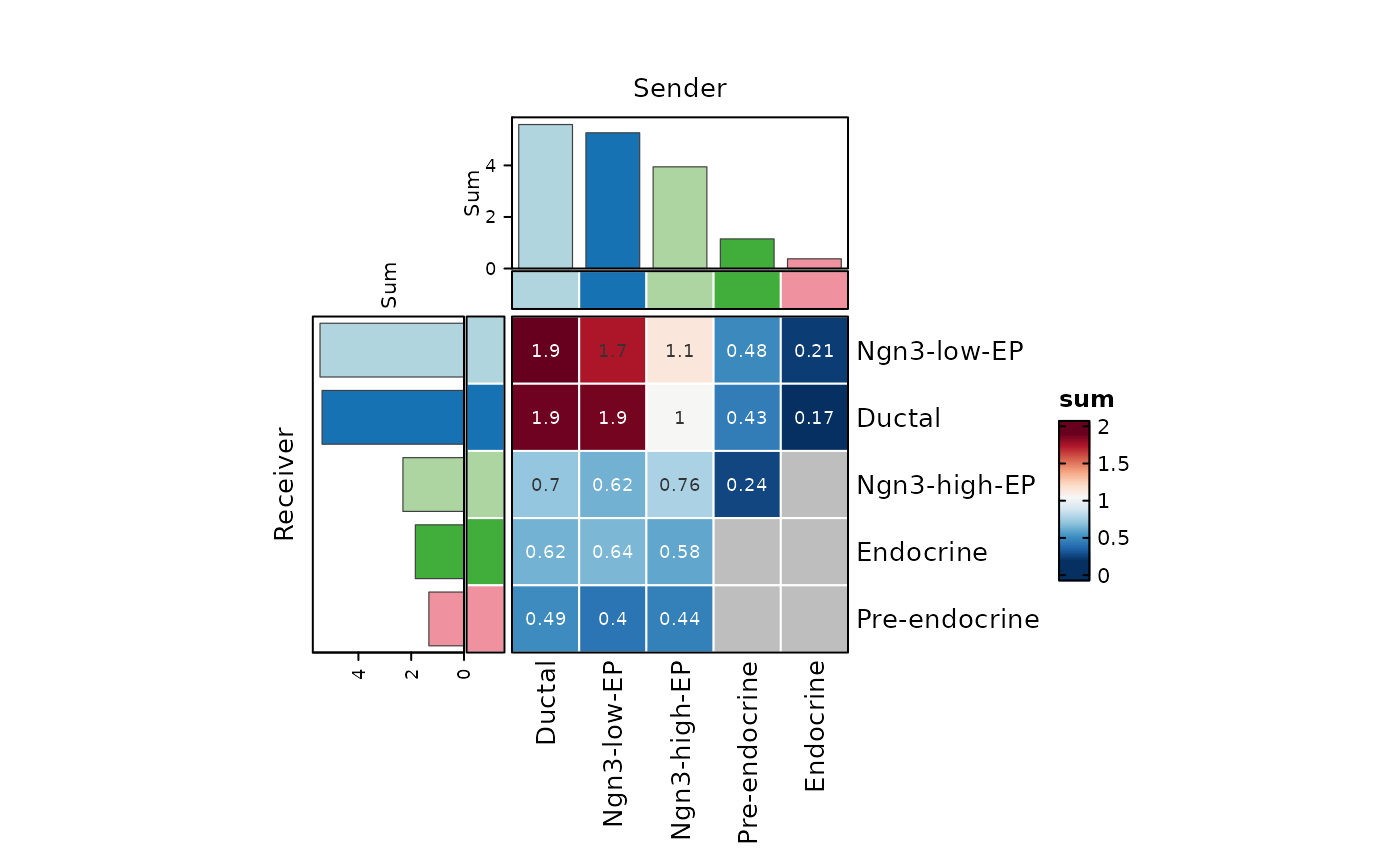 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "dot",
display_by = "interaction",
sender.use = "Ductal",
top_n = 20
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "dot",
display_by = "interaction",
sender.use = "Ductal",
top_n = 20
)
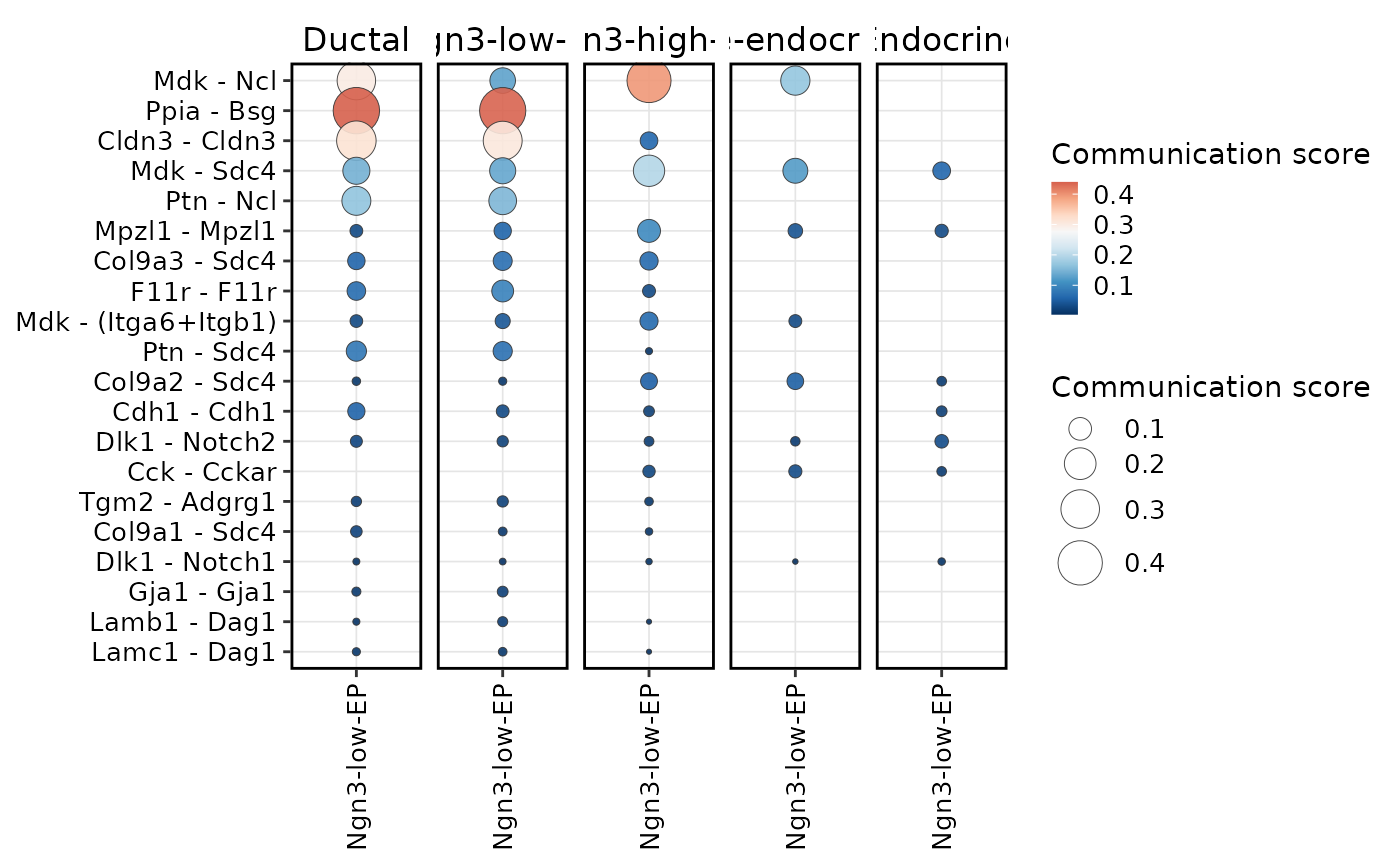
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "dot",
display_by = "interaction",
receiver.use = "Ngn3-low-EP",
top_n = 20
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "dot",
display_by = "interaction",
receiver.use = "Ngn3-low-EP",
top_n = 20
)
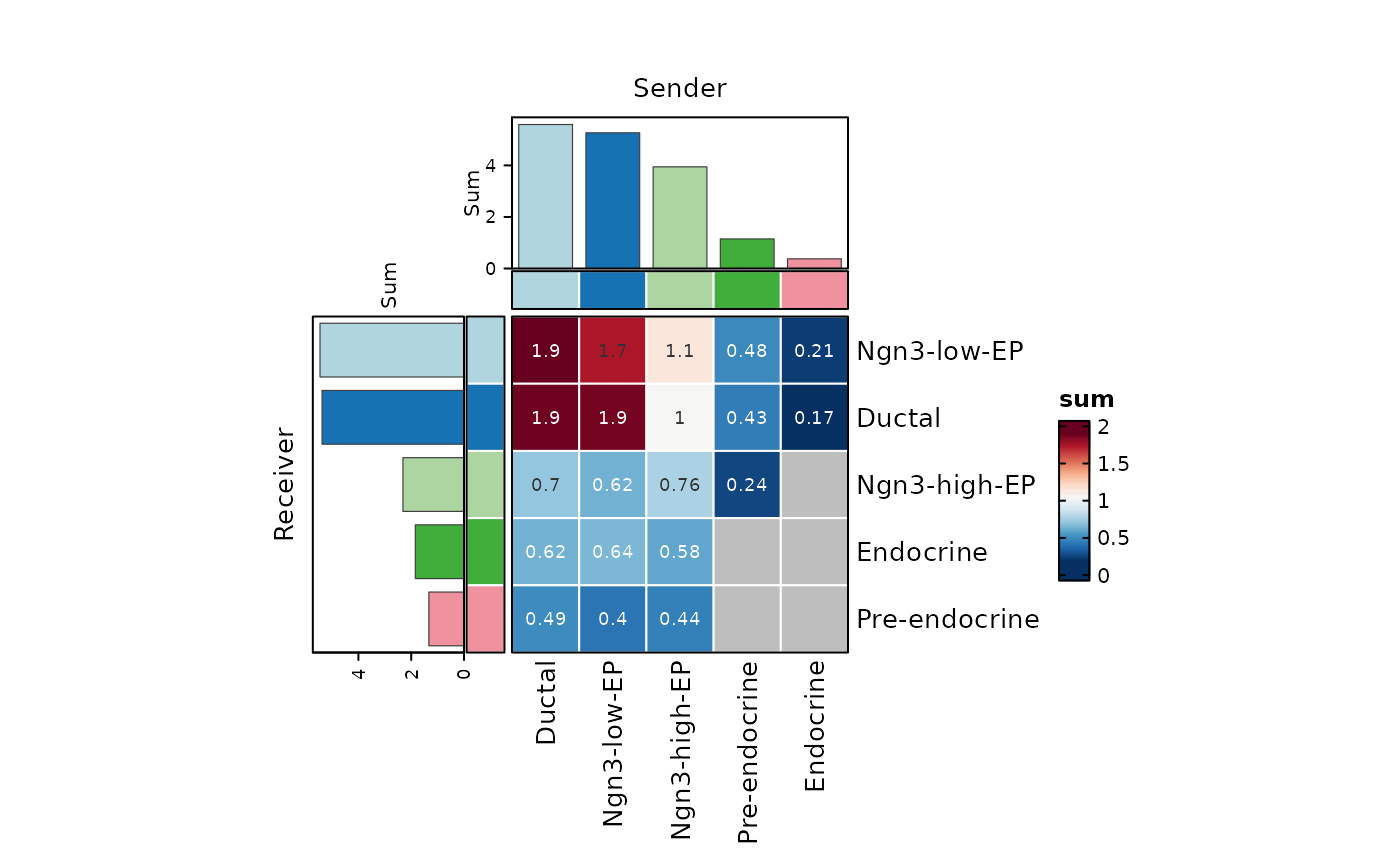
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "aggregation",
top_n = 20
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "aggregation",
top_n = 20
)
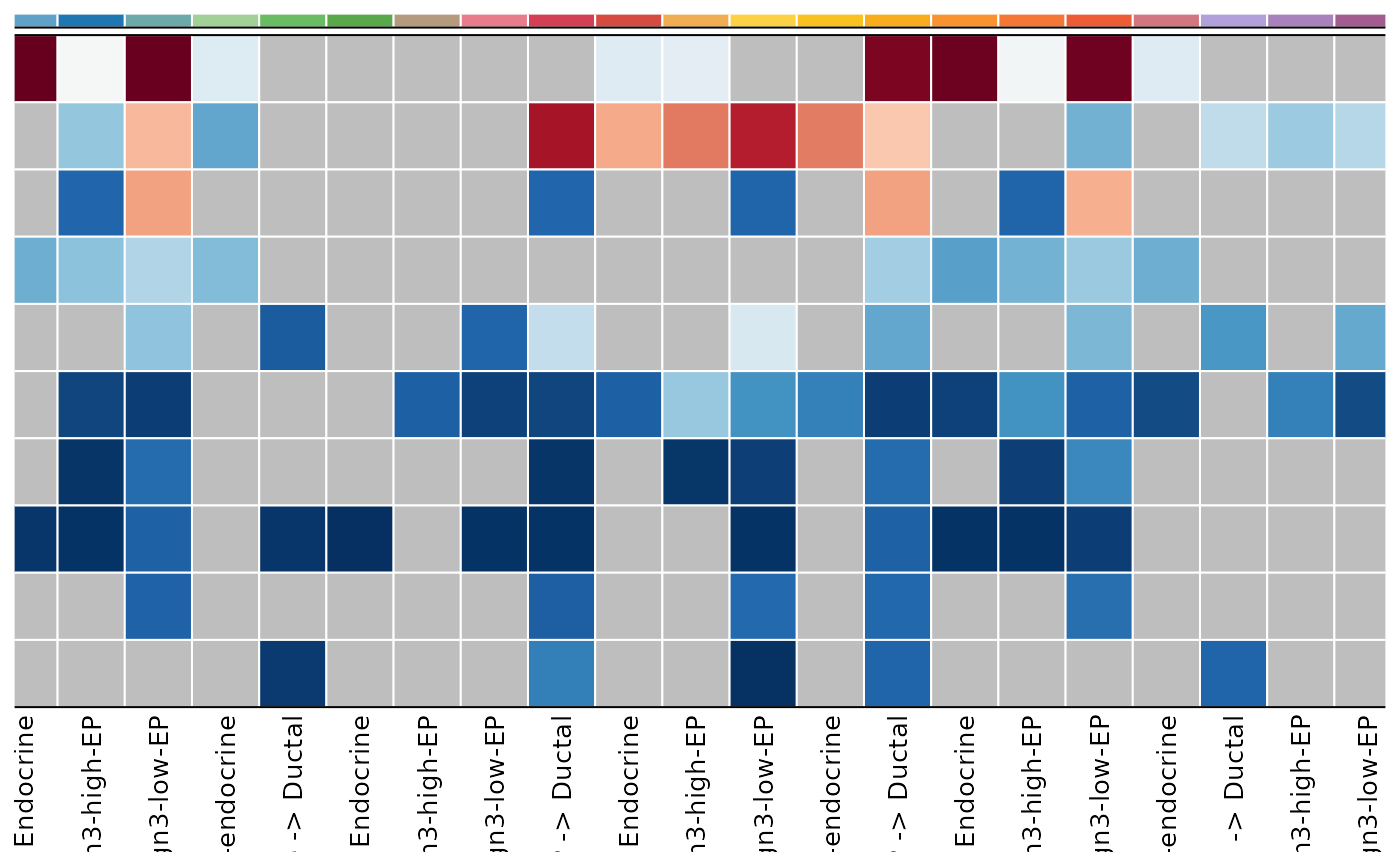
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "interaction",
facet_by = "sender",
top_n = 10
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "interaction",
facet_by = "sender",
top_n = 10
)
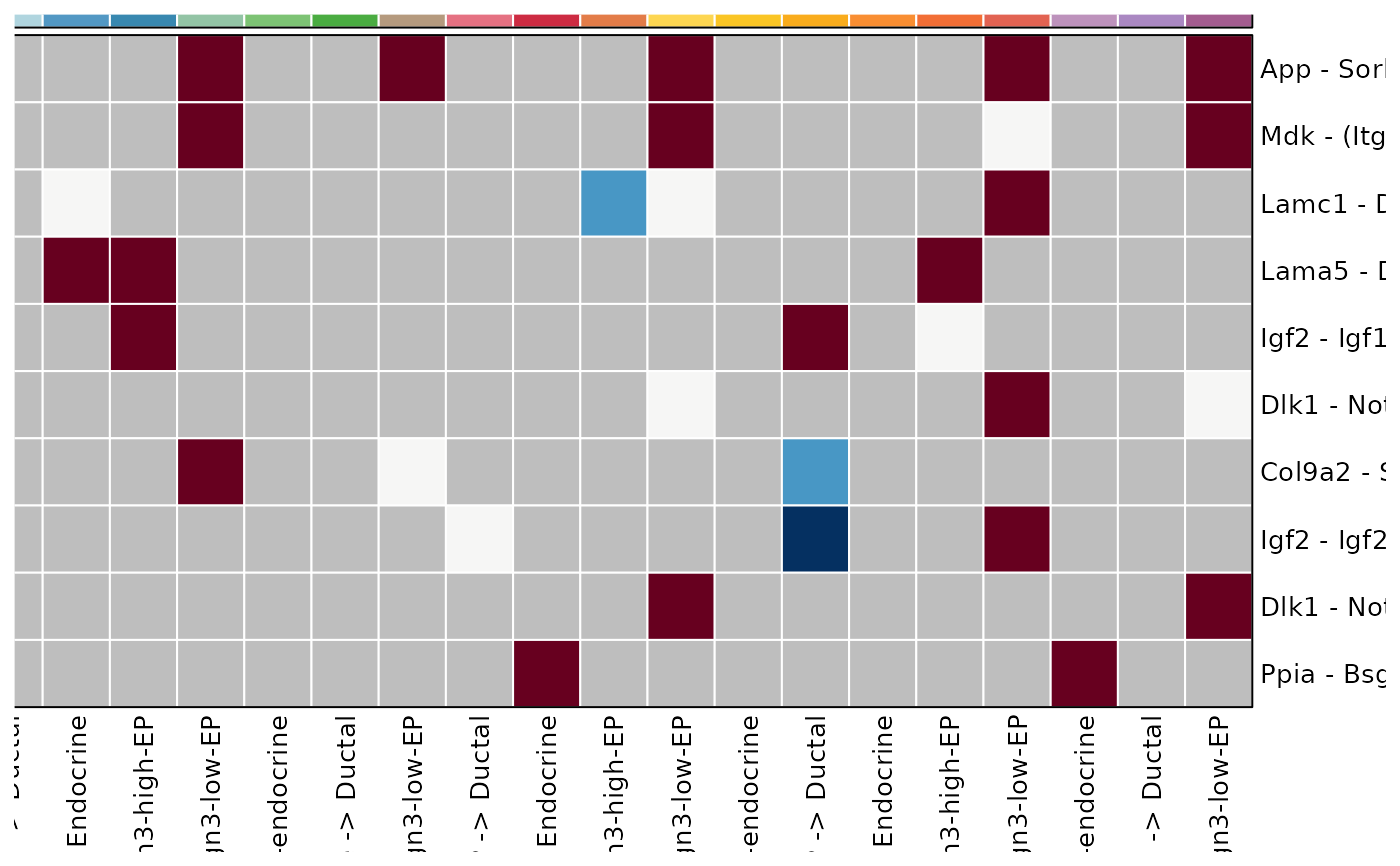
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "interaction",
facet_by = "sender",
color.by = "pvalue",
top_n = 10
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "heatmap",
display_by = "interaction",
facet_by = "sender",
color.by = "pvalue",
top_n = 10
)
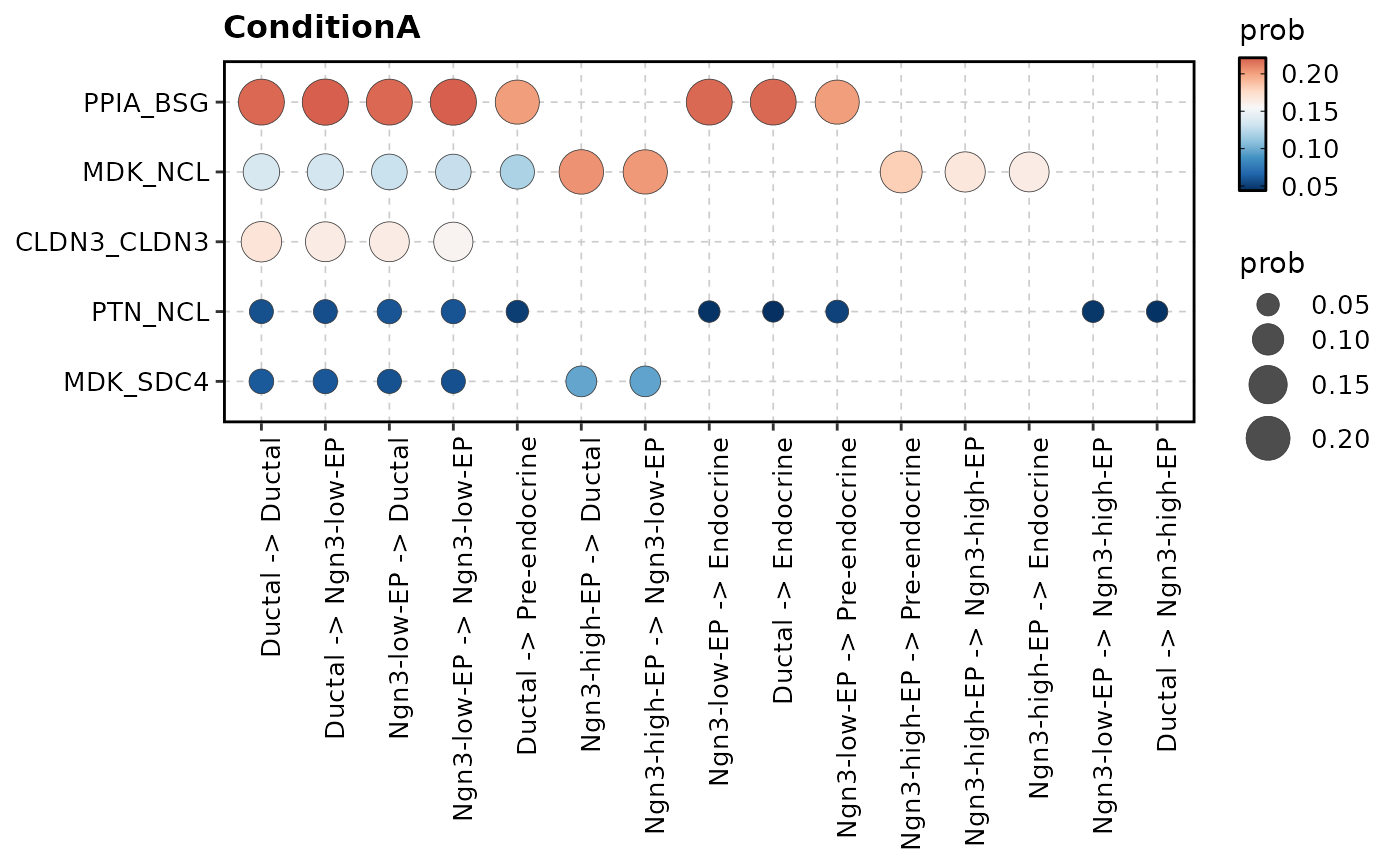
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "bubble",
top_n = 5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "bubble",
top_n = 5
)
 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "bubble",
top_n = 5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "bubble",
top_n = 5
)
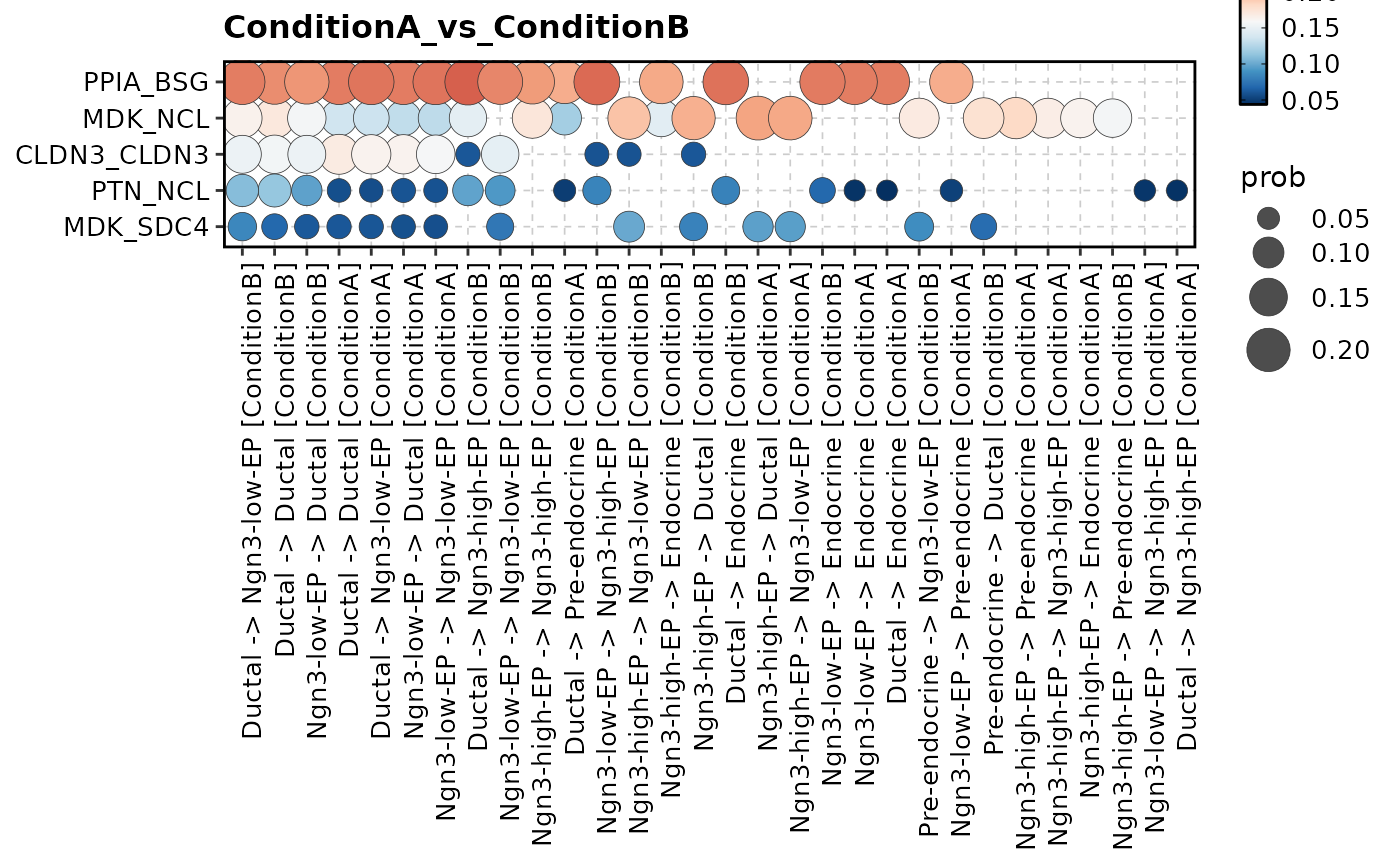 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "role_heatmap",
#' pattern = "outgoing",
width = 0.6,
height = 2.5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "role_heatmap",
#' pattern = "outgoing",
width = 0.6,
height = 2.5
)
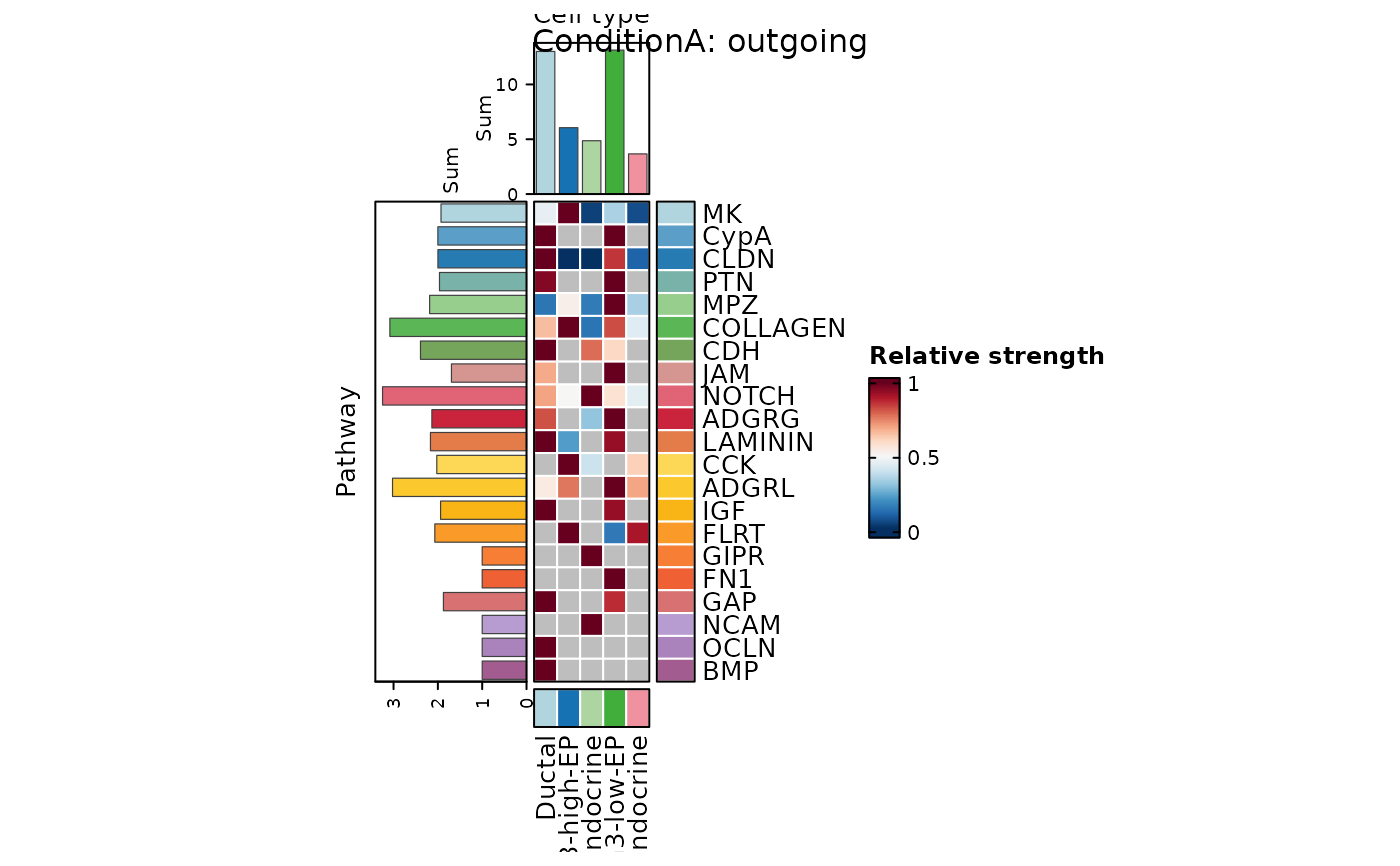 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "role_heatmap",
pattern = "outgoing",
width = 0.6,
height = 2.5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA",
plot_type = "role_heatmap",
pattern = "outgoing",
width = 0.6,
height = 2.5
)
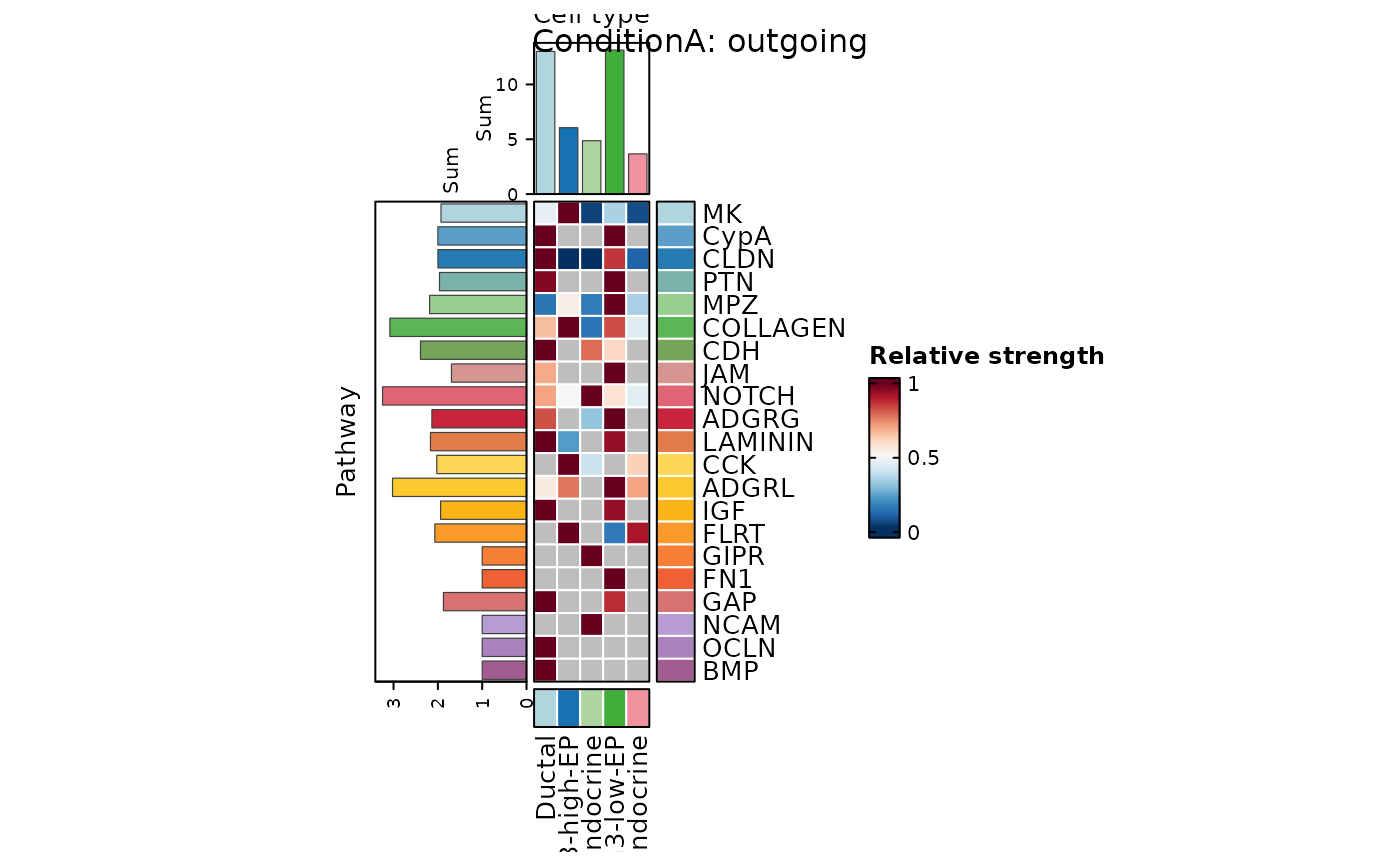 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "role_heatmap",
pattern = "incoming",
palette = "Paired",
width = 0.6,
height = 3.5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "role_heatmap",
pattern = "incoming",
palette = "Paired",
width = 0.6,
height = 3.5
)
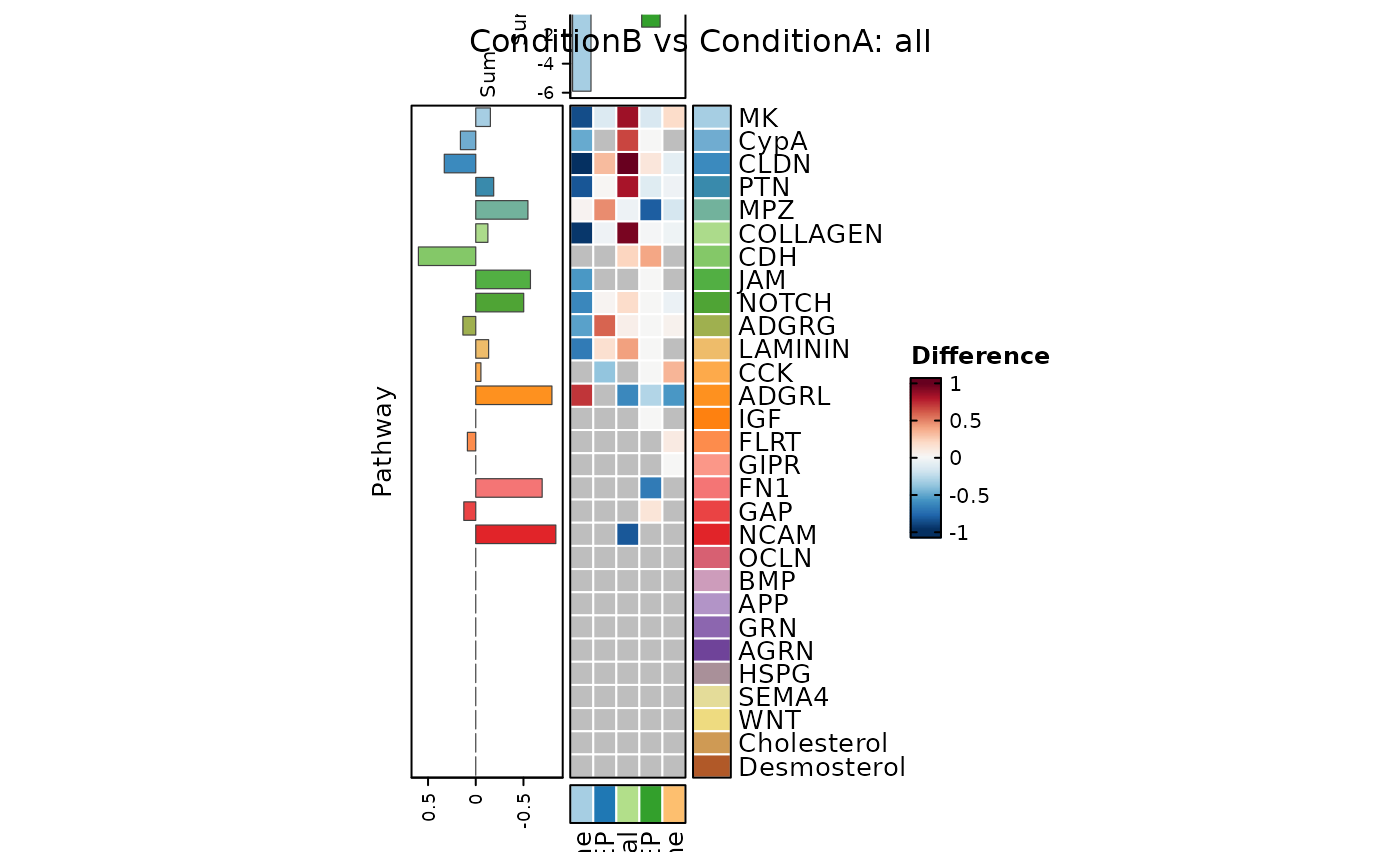
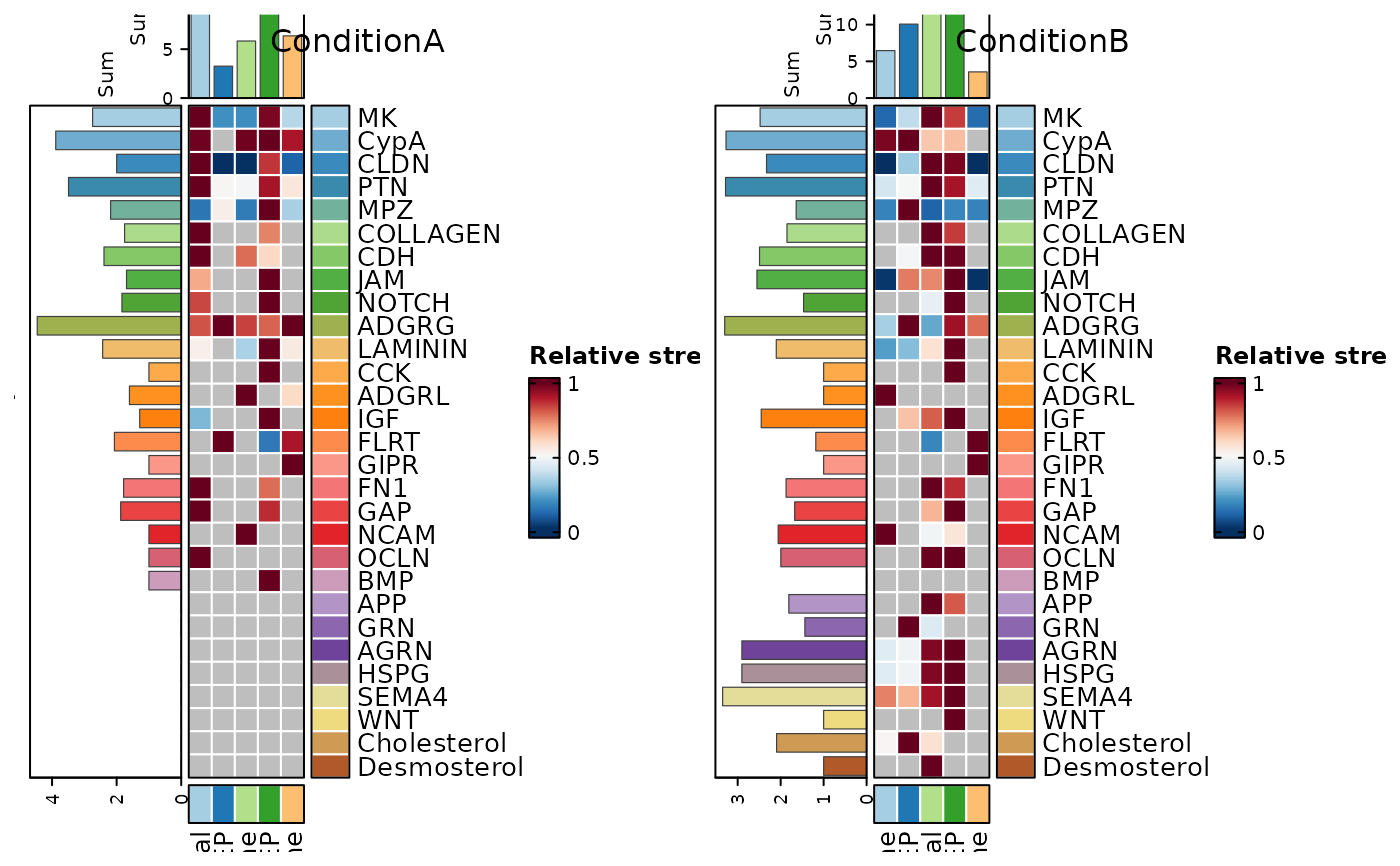 CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "diff_heatmap",
pattern = "all",
palette = "Paired",
top_n = 20,
width = 0.6,
height = 3.5
)
CCCHeatmap(
pancreas_sub,
method = "CellChat",
condition = "ConditionA_vs_ConditionB",
plot_type = "diff_heatmap",
pattern = "all",
palette = "Paired",
top_n = 20,
width = 0.6,
height = 3.5
)