Compute per-cell Local Inverse Simpson's Index (LISI) scores for one or more categorical variables.
This is a clean-room reimplementation of the immunogenomics/LISI.
Arguments
- X
A matrix-like object with cells in rows and embedding/features in columns.
- meta_data
A data frame with one row per cell.
- label_colnames
Character vector of column names in
meta_datato evaluate.- perplexity
Effective neighborhood size. Defaults to
30.- tol
Tolerance used in the binary search for the target perplexity. Defaults to
1e-5.- max_iter
Maximum number of binary-search iterations. Defaults to
50.
References
Korsunsky I, Millard N, Fan J, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nature Methods (2019). https://www.nature.com/articles/s41592-019-0619-0
LISI reference implementation: https://github.com/immunogenomics/LISI
Examples
set.seed(1)
X <- rbind(
matrix(stats::rnorm(100, mean = -1), ncol = 2),
matrix(stats::rnorm(100, mean = 1), ncol = 2)
)
meta_data <- data.frame(
batch = rep(c("A", "B"), each = 50),
group = sample(c("g1", "g2"), 100, replace = TRUE)
)
res <- compute_lisi(
X, meta_data,
c("batch", "group"),
perplexity = 10
)
head(res)
#> batch group
#> 1 1.008491 1.922116
#> 2 1.001277 1.956112
#> 3 1.005979 1.747556
#> 4 1.030285 1.995189
#> 5 1.841283 1.967195
#> 6 1.484192 1.668199
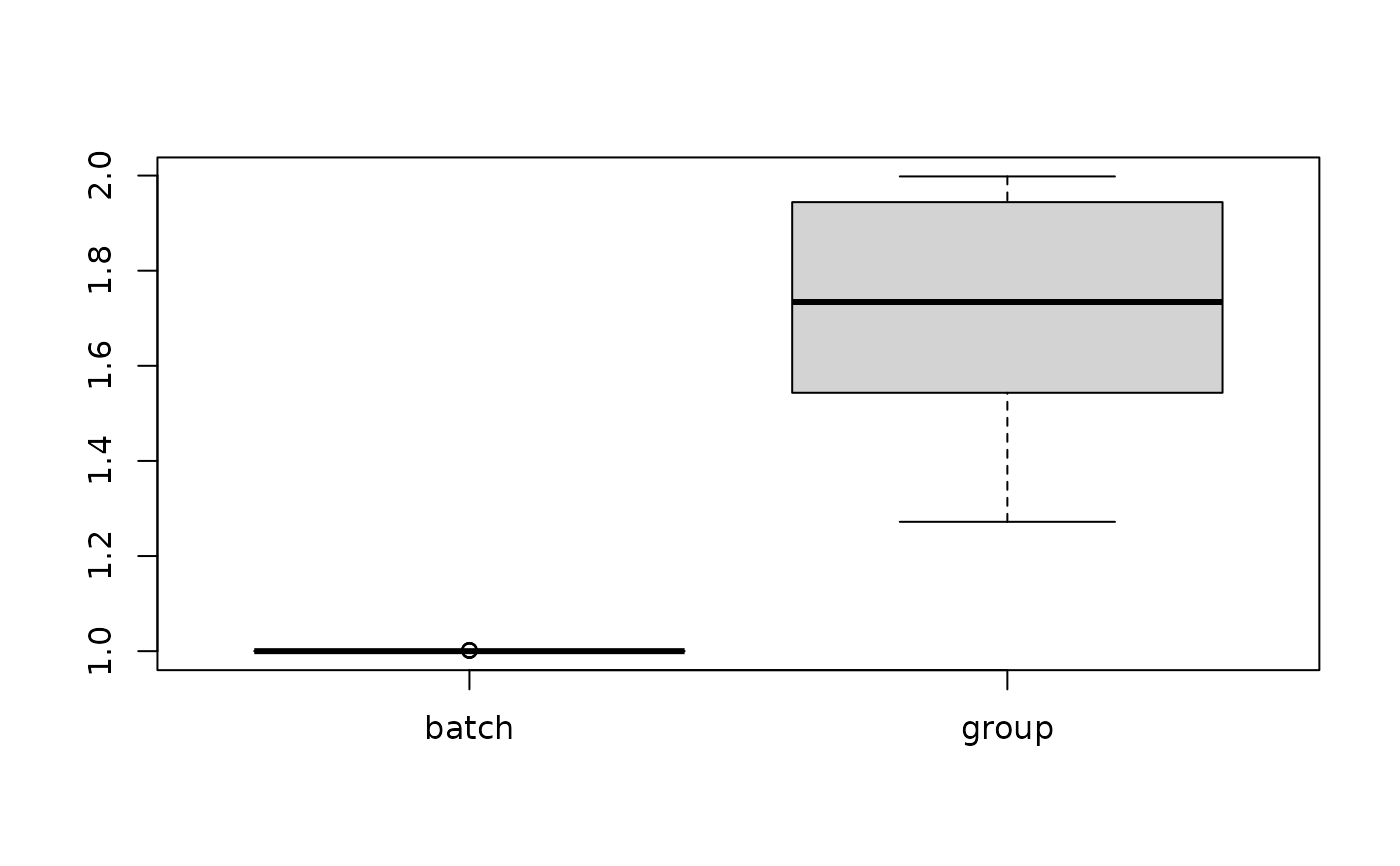
boxplot(res)